Mechanism of the Anticoagulant Activity of Thrombin Mutant W215A/E217A.
Gandhi, P.S., Page, M.J., Chen, Z., Bush-Pelc, L., Di Cera, E.(2009) J Biological Chem 284: 24098-24105
- PubMed: 19586901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.025403
- Primary Citation Related Structures:
3HK3, 3HK6, 3HKI, 3HKJ - PubMed Abstract:
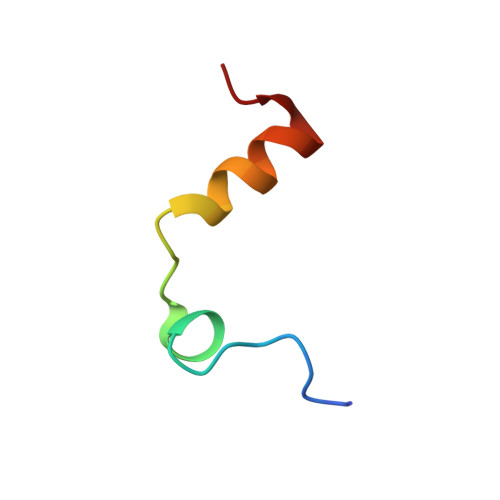
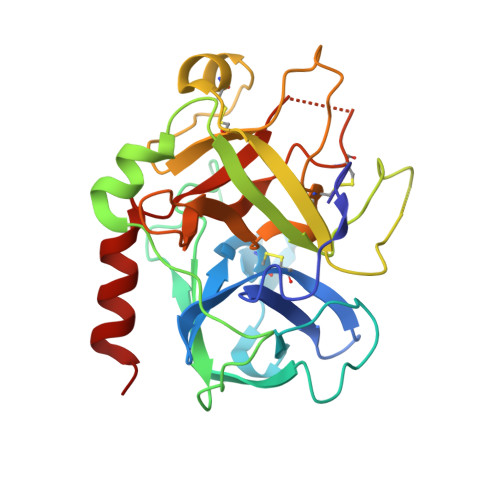
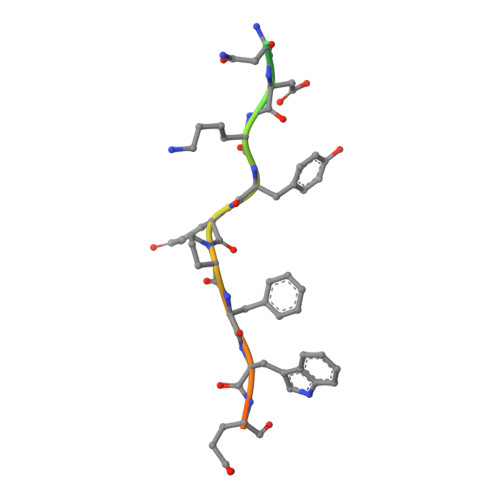
The thrombin mutant W215A/E217A (WE) is a potent anticoagulant both in vitro and in vivo. Previous x-ray structural studies have shown that WE assumes a partially collapsed conformation that is similar to the inactive E* form, which explains its drastically reduced activity toward substrate. Whether this collapsed conformation is genuine, rather than the result of crystal packing or the mutation introduced in the critical 215-217 beta-strand, and whether binding of thrombomodulin to exosite I can allosterically shift the E* form to the active E form to restore activity toward protein C are issues of considerable mechanistic importance to improve the design of an anticoagulant thrombin mutant for therapeutic applications. Here we present four crystal structures of WE in the human and murine forms that confirm the collapsed conformation reported previously under different experimental conditions and crystal packing. We also present structures of human and murine WE bound to exosite I with a fragment of the platelet receptor PAR1, which is unable to shift WE to the E form. These structural findings, along with kinetic and calorimetry data, indicate that WE is strongly stabilized in the E* form and explain why binding of ligands to exosite I has only a modest effect on the E*-E equilibrium for this mutant. The E* --> E transition requires the combined binding of thrombomodulin and protein C and restores activity of the mutant WE in the anticoagulant pathway.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110, USA.
Organizational Affiliation: