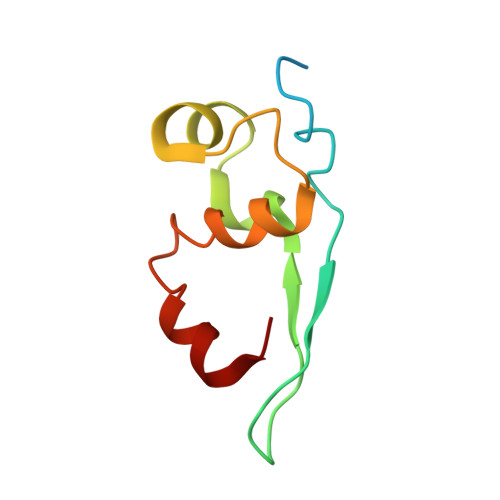
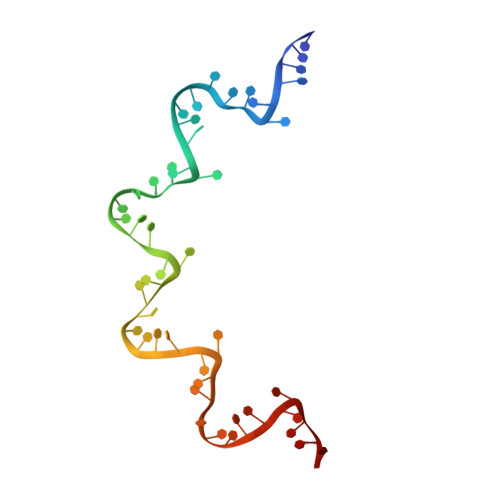
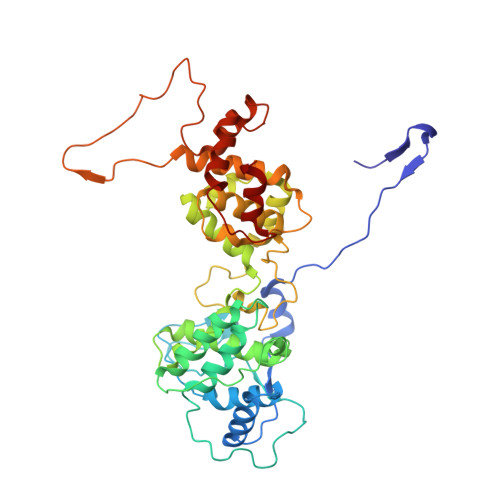
Structure of the vesicular stomatitis virus nucleocapsid in complex with the nucleocapsid-binding domain of the small polymerase cofactor, P.
Green, T.J., Luo, M.(2009) Proc Natl Acad Sci U S A 106: 11713-11718
- PubMed: 19571006
- DOI: https://doi.org/10.1073/pnas.0903228106
- Primary Citation of Related Structures:
3HHW, 3HHZ - PubMed Abstract:
The negative-strand RNA viruses (NSRVs) are unique because their nucleocapsid, not the naked RNA, is the active template for transcription and replication. The viral polymerase of nonsegmented NSRVs contains a large polymerase catalytic subunit (L) and a nonenzymatic cofactor, the phosphoprotein (P). Insight into how P delivers the polymerase complex to the nucleocapsid has long been pursued by reverse genetics and biochemical approaches. Here, we present the X-ray crystal structure of the C-terminal domain of P of vesicular stomatitis virus, a prototypic nonsegmented NSRV, bound to nucleocapsid-like particles. P binds primarily to the C-terminal lobe of 2 adjacent N proteins within the nucleocapsid. This binding mode is exclusive to the nucleocapsid, not the nucleocapsid (N) protein in other existing forms. Localization of phosphorylation sites within P and their proximity to the RNA cavity give insight into how the L protein might be oriented to access the RNA template.
- Department of Microbiology, University of Alabama School of Medicine, Birmingham, AL 35294, USA.
Organizational Affiliation: