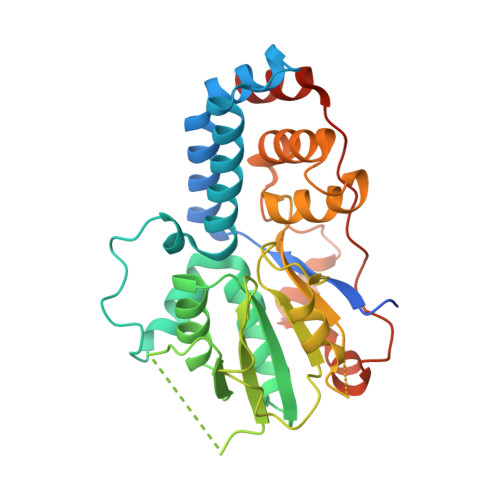
Structural and Functional Studies of the Yeast Class II Hda1 Histone Deacetylase Complex.
Lee, J.H., Maskos, K., Huber, R.(2009) J Mol Biology 391: 744-757
- PubMed: 19573535
- DOI: https://doi.org/10.1016/j.jmb.2009.06.059
- Primary Citation of Related Structures:
3HGQ, 3HGT - PubMed Abstract:
Yeast class II Hda1 histone deacetylase (HDAC) complex is an H2B- and H3-specific HDAC in Saccharomyces cerevisiae consisting of three previously identified subunits, the catalytic subunit scHda1p and two non-catalytic structural subunits scHda2p and scHda3p. We co-expressed and co-purified recombinant yeast class II HDAC complex from bacteria as a functionally active and trichostatin-A-sensitive hetero-tetrameric complex. According to an extensive analysis of domain organization and interaction of all subunits (or domains), the N-terminal domain of scHda1p associates through the C-terminal coiled-coil domains (CCDs) of the scHda2p-scHda3p sub-complex, yielding a truncated scHda1pHDAC-scHda2pCCD2-scHda3pCCD3 complex with indistinguishable deacetylase activity compared to the full-length complex in vitro. We characterized the interaction of the HDAC complex with either single-stranded or double-stranded DNA and identified the N-terminal halves of scHda2p and scHda3p as binding modules. A high-resolution structure of the scHda3p DNA-binding domain by X-ray crystallography is presented. The crystal structure shows an unanticipated structural homology with the C-terminal helicase lobes of SWI2/SNF2 chromatin-remodeling domains of the Rad54 family enzymes. DNA binding is unspecific for nucleotide sequence and structure, similar to the Rad54 enzymes in vitro. Our structural and functional analyses of the budding yeast class II Hda1 HDAC complex provide insight into DNA recognition and deacetylation of histones in nucleosomes.
- Max-Planck-Institute of Biochemistry, Martinsried, Germany. lee@biochem.mpg.de
Organizational Affiliation: