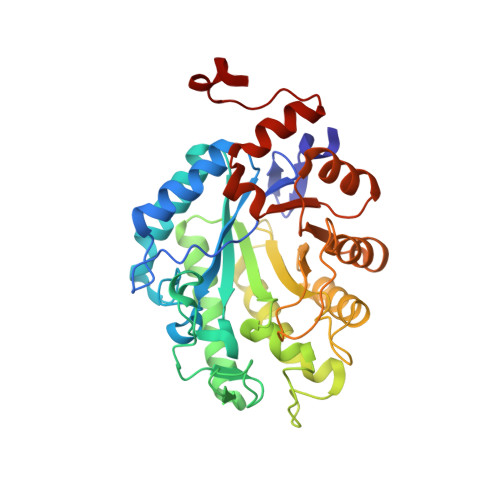
Crystal structure of a thermostable old yellow enzyme from Thermus scotoductus SA-01
Opperman, D.J., Sewell, B.T., Litthauer, D., Isupov, M.N., Littlechild, J.A., van Heerden, E.(2010) Biochem Biophys Res Commun 393: 426-431
- PubMed: 20138824 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2010.02.011
- Primary Citation Related Structures:
3HF3, 3HGJ - PubMed Abstract:
Recent characterization of the chromate reductase (CrS) from the thermophile Thermus scotoductus SA-01 revealed this enzyme to be related to the Old Yellow Enzyme (OYE) family. Here, we report the structure of a thermostable OYE homolog in its holoform at 2.2A as well as its complex with p-hydroxybenzaldehyde (pHBA). The enzyme crystallized as octamers with the monomers showing a classical TIM barrel fold which upon dimerization yields the biologically active form of the protein. A sulfate ion is bound above the si-side of the non-covalently bound FMN cofactor in the oxidized solved structure but is displaced upon pHBA binding. The active-site architecture is highly conserved as with other members of this enzyme family. The pHBA in the CrS complex is positioned by hydrogen bonding to the two conserved catalytic-site histidines. The most prominent structural difference between CrS and other OYE homologs is the size of the "capping domain". Thermostabilization of the enzyme is achieved in part through increased proline content within loops and turns as well as increased intersubunit interactions through hydrogen bonding and complex salt bridge networks. CrS is able to reduce the C=C bonds of alpha,beta-unsaturated carbonyl compounds with a preference towards cyclic substrates however no activity was observed towards beta-substituted substrates. Mutational studies have confirmed the role of Tyr177 as the proposed proton donor although reduction could still occur at a reduced rate when this residue was mutated to phenylalanine.
- Department of Microbial, Biochemical and Food Biotechnology, BioPAD Metagenomics Platform, University of the Free State, Bloemfontein 9300, South Africa.
Organizational Affiliation: