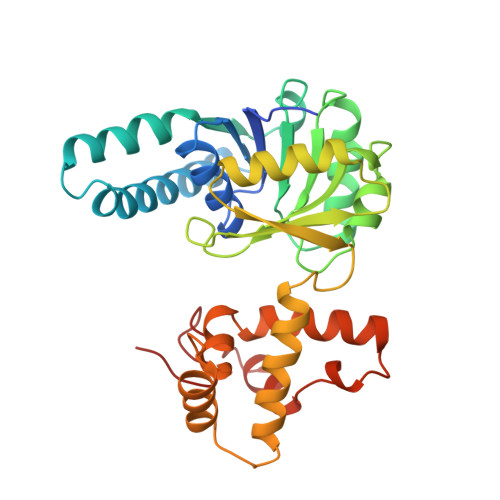
Pig heart short chain L-3-hydroxyacyl-CoA dehydrogenase revisited: sequence analysis and crystal structure determination.
Barycki, J.J., O'Brien, L.K., Birktoft, J.J., Strauss, A.W., Banaszak, L.J.(1999) Protein Sci 8: 2010-2018
- PubMed: 10548046 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.8.10.2010
- Primary Citation Related Structures:
3HDH - PubMed Abstract:
Short chain L-3-hydroxyacyl CoA dehydrogenase (SCHAD) is a soluble dimeric enzyme critical for oxidative metabolism of fatty acids. Its primary sequence has been reported to be conserved across numerous tissues and species with the notable exception of the pig heart homologue. Preliminary efforts to solve the crystal structure of the dimeric pig heart SCHAD suggested the unprecedented occurrence of three enzyme subunits within the asymmetric unit, a phenomenon that was thought to have hampered refinement of the initial chain tracing. The recently solved crystal coordinates of human heart SCHAD facilitated a molecular replacement solution to the pig heart SCHAD data. Refinement of the model, in conjunction with the nucleotide sequence for pig heart SCHAD determined in this paper, has demonstrated that the previously published pig heart SCHAD sequence was incorrect. Presented here are the corrected amino acid sequence and the high resolution crystal structure determined for pig heart SCHAD complexed with its NAD+ cofactor (2.8 A; R(cryst) = 22.4%, R(free) = 28.8%). In addition, the peculiar phenomenon of a dimeric enzyme crystallizing with three subunits contained in the asymmetric unit is described.
- Department of Biochemistry, Molecular Biology, and Biophysics, University of Minnesota, Minneapolis 55455, USA.
Organizational Affiliation: