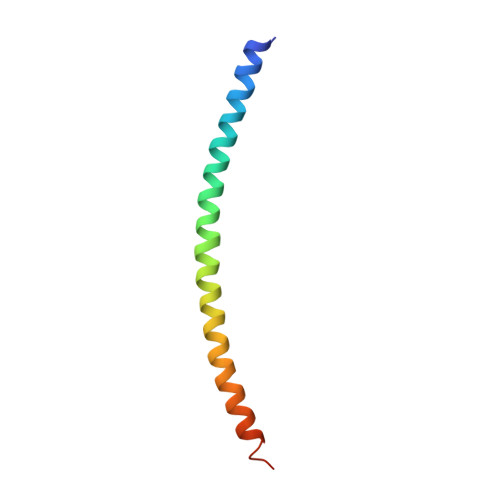
Helical extension of the neuronal SNARE complex into the membrane
Stein, A., Weber, G., Wahl, M.C., Jahn, R.(2009) Nature 460: 525-528
- PubMed: 19571812 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08156
- Primary Citation Related Structures:
3HD7, 3IPD - PubMed Abstract:
Neurotransmission relies on synaptic vesicles fusing with the membrane of nerve cells to release their neurotransmitter content into the synaptic cleft, a process requiring the assembly of several members of the SNARE (soluble N-ethylmaleimide-sensitive factor attachment protein receptor) family. SNAREs represent an evolutionarily conserved protein family that mediates membrane fusion in the secretory and endocytic pathways of eukaryotic cells. On membrane contact, these proteins assemble in trans between the membranes as a bundle of four alpha-helices, with the energy released during assembly being thought to drive fusion. However, it is unclear how the energy is transferred to the membranes and whether assembly is conformationally linked to fusion. Here, we report the X-ray structure of the neuronal SNARE complex, consisting of rat syntaxin 1A, SNAP-25 and synaptobrevin 2, with the carboxy-terminal linkers and transmembrane regions at 3.4 A resolution. The structure shows that assembly proceeds beyond the already known core SNARE complex, resulting in a continuous helical bundle that is further stabilized by side-chain interactions in the linker region. Our results suggest that the final phase of SNARE assembly is directly coupled to membrane merger.
- Department of Neurobiology, Max Planck Institute for Biophysical Chemistry, 37077 Göttingen, Germany.
Organizational Affiliation: