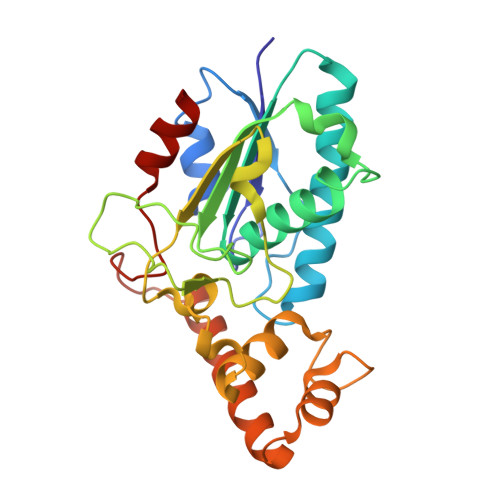
Mycobacteriophage Lysin B is a novel mycolylarabinogalactan esterase
Payne, K., Sun, Q., Sacchettini, J., Hatfull, G.F.(2009) Mol Microbiol 73: 367-381
- PubMed: 19555454 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1365-2958.2009.06775.x
- Primary Citation Related Structures:
3HC7 - PubMed Abstract:
Mycobacteriophages encounter a unique problem among phages of Gram-positive bacteria, in that lysis must not only degrade the peptidoglycan layer but also circumvent a mycolic acid-rich outer membrane covalently attached to the arabinogalactan-peptidoglycan complex. Mycobacteriophages accomplish this by producing two lysis enzymes, Lysin A (LysA) that hydrolyses peptidoglycan, and Lysin B (LysB), a novel mycolylarabinogalactan esterase, that cleaves the mycolylarabinogalactan bond to release free mycolic acids. The D29 LysB structure shows an alpha/beta hydrolase organization with a catalytic triad common to cutinases, but which contains an additional four-helix domain implicated in the binding of lipid substrates. Whereas LysA is essential for mycobacterial lysis, a Giles DeltalysB mutant mycobacteriophage is viable, but defective in the normal timing, progression and completion of host cell lysis. We propose that LysB facilitates lysis by compromising the integrity of the mycobacterial outer membrane linkage to the arabinogalactan-peptidoglycan layer.
- Department of Biological Sciences, University of Pittsburgh, Pittsburgh, PA 15260, USA.
Organizational Affiliation: