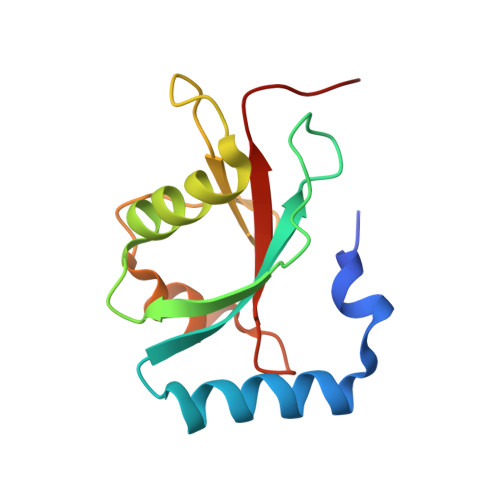
Trypanosoma brucei ATG8: structural insights into autophagic-like mechanisms in protozoa.
Koopmann, R., Muhammad, K., Perbandt, M., Betzel, C., Duszenko, M.(2009) Autophagy 5: 1085-1091
- PubMed: 19736525 Search on PubMed
- DOI: https://doi.org/10.4161/auto.5.8.9611
- Primary Citation Related Structures:
3H9D - PubMed Abstract:
Bioinformatic searches of genome databases revealed that the number of autophagy-related genes (ATG) is considerably lower in trypanosomes than in higher eukaryotes and even in yeast. This raises the question of whether autophagy in this protozoan parasite is more primitive and represents a rudimentary paradigm due to its early branching off the evolutionary tree. We here present the crystal structure of TbATG8B. This molecule (MW 13.7 kDa) belongs to the ubiquitin-like proteins showing the typical ubiquitin fold and strong sequence homology to LC3, the human homologue. Due to its characteristic folding, it should readily bind to TbATG4.1 for being processed. This presumption was tested by molecular modeling approaches, docking TbATG8B to a homology model of TbATG4.1. Although exchanges of several amino acids are evident from sequence comparisons, the overall structure seems very much alike and the necessary catalytic triad (C-D-H) is well conserved in TbATG4.1. Thus membrane formation during appearance of the autophagic bodies seems very similar in trypanosomes and their higher eukaryotic counterpart.
- Interfaculty Institute for Biochemistry, University of Tübingen, Tübingen, Germany.
Organizational Affiliation: