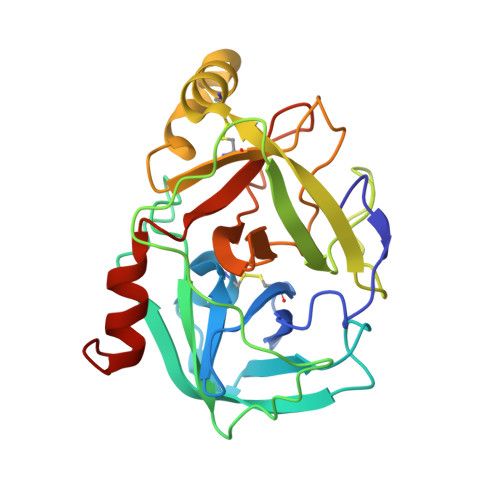
Structural mechanisms of inactivation in scabies mite serine protease paralogues.
Fischer, K., Langendorf, C.G., Irving, J.A., Reynolds, S., Willis, C., Beckham, S., Law, R.H., Yang, S., Bashtannyk-Puhalovich, T.A., McGowan, S., Whisstock, J.C., Pike, R.N., Kemp, D.J., Buckle, A.M.(2009) J Mol Biology 390: 635-645
- PubMed: 19427318 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.04.082
- Primary Citation Related Structures:
3H7O, 3H7T - PubMed Abstract:
The scabies mite (Sarcoptes scabiei) is a parasite responsible for major morbidity in disadvantaged communities and immuno-compromised patients worldwide. In addition to the physical discomfort caused by the disease, scabies infestations facilitate infection by Streptococcal species via skin lesions, resulting in a high prevalence of rheumatic fever/heart disease in affected communities. The scabies mite produces 33 proteins that are closely related to those in the dust mite group 3 allergen and belong to the S1-like protease family (chymotrypsin-like). However, all but one of these molecules contain mutations in the conserved active-site catalytic triad that are predicted to render them catalytically inactive. These molecules are thus termed scabies mite inactivated protease paralogues (SMIPPs). The precise function of SMIPPs is unclear; however, it has been suggested that these proteins might function by binding and protecting target substrates from cleavage by host immune proteases, thus preventing the host from mounting an effective immune challenge. In order to begin to understand the structural basis for SMIPP function, we solved the crystal structures of SMIPP-S-I1 and SMIPP-S-D1 at 1.85 A and 2.0 A resolution, respectively. Both structures adopt the characteristic serine protease fold, albeit with large structural variations over much of the molecule. In both structures, mutations in the catalytic triad together with occlusion of the S1 subsite by a conserved Tyr200 residue is predicted to block substrate ingress. Accordingly, we show that both proteases lack catalytic function. Attempts to restore function (via site-directed mutagenesis of catalytic residues as well as Tyr200) were unsuccessful. Taken together, these data suggest that SMIPPs have lost the ability to bind substrates in a classical "canonical" fashion, and instead have evolved alternative functions in the lifecycle of the scabies mite.
- Scabies Laboratory, Queensland Institute of Medical Research, Brisbane, Australia. Katja.Fischer@qimr.edu.au
Organizational Affiliation: