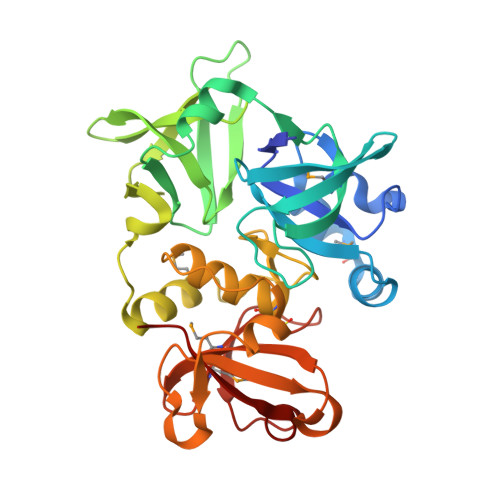
Structure of the gamma-D-glutamyl-L-diamino acid endopeptidase YkfC from Bacillus cereus in complex with L-Ala-[gamma]-D-Glu: insights into substrate recognition by NlpC/P60 cysteine peptidases
Xu, Q., Abdubek, P., Astakhova, T., Axelrod, H.L., Bakolitsa, C., Cai, X., Carlton, D., Chen, C., Chiu, H.J., Chiu, M., Clayton, T., Das, D., Deller, M.C., Duan, L., Ellrott, K., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Lam, W.W., Marciano, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Wooten, T., Yeh, A., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1354-1364
- PubMed: 20944232 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110021214
- Primary Citation Related Structures:
3H41 - PubMed Abstract:
Dipeptidyl-peptidase VI from Bacillus sphaericus and YkfC from Bacillus subtilis have both previously been characterized as highly specific γ-D-glutamyl-L-diamino acid endopeptidases. The crystal structure of a YkfC ortholog from Bacillus cereus (BcYkfC) at 1.8 Å resolution revealed that it contains two N-terminal bacterial SH3 (SH3b) domains in addition to the C-terminal catalytic NlpC/P60 domain that is ubiquitous in the very large family of cell-wall-related cysteine peptidases. A bound reaction product (L-Ala-γ-D-Glu) enabled the identification of conserved sequence and structural signatures for recognition of L-Ala and γ-D-Glu and, therefore, provides a clear framework for understanding the substrate specificity observed in dipeptidyl-peptidase VI, YkfC and other NlpC/P60 domains in general. The first SH3b domain plays an important role in defining substrate specificity by contributing to the formation of the active site, such that only murein peptides with a free N-terminal alanine are allowed. A conserved tyrosine in the SH3b domain of the YkfC subfamily is correlated with the presence of a conserved acidic residue in the NlpC/P60 domain and both residues interact with the free amine group of the alanine. This structural feature allows the definition of a subfamily of NlpC/P60 enzymes with the same N-terminal substrate requirements, including a previously characterized cyanobacterial L-alanine-γ-D-glutamate endopeptidase that contains the two key components (an NlpC/P60 domain attached to an SH3b domain) for assembly of a YkfC-like active site.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: