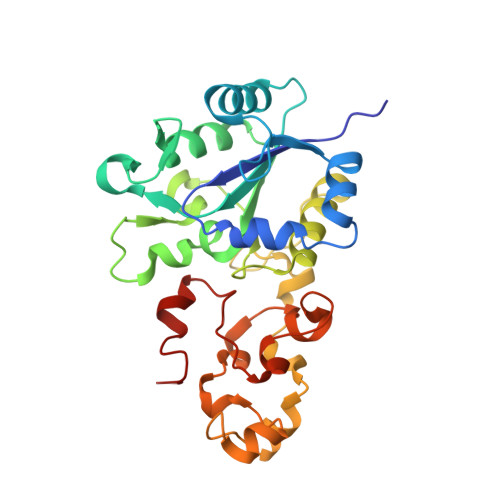
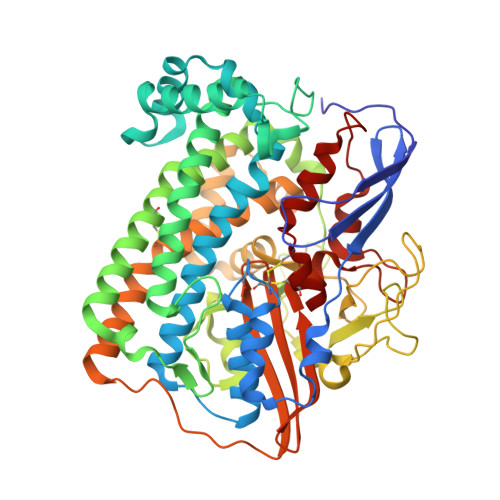
Introduction of methionines in the gas channel makes [NiFe] hydrogenase aero-tolerant
Dementin, S., Leroux, F., Cournac, L., de Lacey, A.L., Volbeda, A., Leger, C., Burlat, B., Martinez, N., Champ, S., Martin, L., Sanganas, O., Haumann, M., Fernandez, V.M., Guigliarelli, B., Fontecilla-Camps, J.C., Rousset, M.(2009) J Am Chem Soc 131: 10156-10164
- PubMed: 19580279 Search on PubMed
- DOI: https://doi.org/10.1021/ja9018258
- Primary Citation Related Structures:
3H3X - PubMed Abstract:
Hydrogenases catalyze the conversion between 2H(+) + 2e(-) and H(2)(1). Most of these enzymes are inhibited by O(2), which represents a major drawback for their use in biotechnological applications. Improving hydrogenase O(2) tolerance is therefore a major contemporary challenge to allow the implementation of a sustainable hydrogen economy. We succeeded in improving O(2) tolerance, which we define here as the ability of the enzyme to resist for several minutes to O(2) exposure, by substituting with methionines small hydrophobic residues strongly conserved in the gas channel. Remarkably, the mutated enzymes remained active in the presence of an O(2) concentration close to that found in aerobic solutions in equilibrium with air, while the wild type enzyme is inhibited in a few seconds. Crystallographic and spectroscopic studies showed that the structure and the chemistry at the active site are not affected by the mutations. Kinetic studies demonstrated that the inactivation is slower and reactivation faster in these mutants. We propose that in addition to restricting O(2) diffusion to the active site of the enzyme, methionine may also interact with bound peroxide and provide an assisted escape route for H(2)O(2) toward the gas channel. These results show for the first time that it is possible to improve O(2)-tolerance of [NiFe] hydrogenases, making possible the development of biohydrogen production systems.
- CNRS, Bioénergétique et Ingénierie des Protéines, IMM, 31 Chemin Joseph Aiguier, 13402 Marseille Cedex 20, France.
Organizational Affiliation: