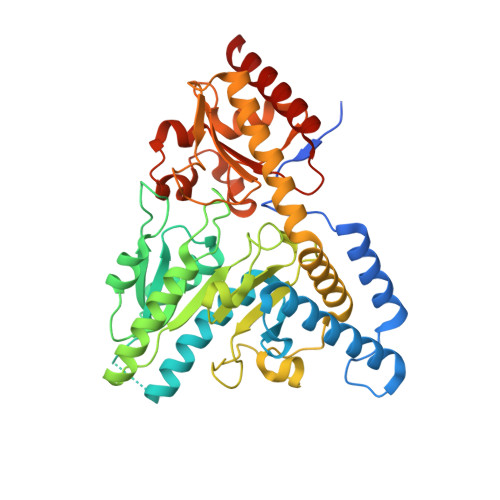
Biochemical discrimination between selenium and sulfur 1: a single residue provides selenium specificity to human selenocysteine lyase.
Collins, R., Johansson, A.L., Karlberg, T., Markova, N., van den Berg, S., Olesen, K., Hammarstrom, M., Flores, A., Schuler, H., Schiavone, L.H., Brzezinski, P., Arner, E.S., Hogbom, M.(2012) PLoS One 7: e30581-e30581
- PubMed: 22295093
- DOI: https://doi.org/10.1371/journal.pone.0030581
- Primary Citation Related Structures:
3GZC, 3GZD - PubMed Abstract:
Selenium and sulfur are two closely related basic elements utilized in nature for a vast array of biochemical reactions. While toxic at higher concentrations, selenium is an essential trace element incorporated into selenoproteins as selenocysteine (Sec), the selenium analogue of cysteine (Cys). Sec lyases (SCLs) and Cys desulfurases (CDs) catalyze the removal of selenium or sulfur from Sec or Cys and generally act on both substrates. In contrast, human SCL (hSCL) is specific for Sec although the only difference between Sec and Cys is the identity of a single atom. The chemical basis of this selenium-over-sulfur discrimination is not understood. Here we describe the X-ray crystal structure of hSCL and identify Asp146 as the key residue that provides the Sec specificity. A D146K variant resulted in loss of Sec specificity and appearance of CD activity. A dynamic active site segment also provides the structural prerequisites for direct product delivery of selenide produced by Sec cleavage, thus avoiding release of reactive selenide species into the cell. We thus here define a molecular determinant for enzymatic specificity discrimination between a single selenium versus sulfur atom, elements with very similar chemical properties. Our findings thus provide molecular insights into a key level of control in human selenium and selenoprotein turnover and metabolism.
- Structural Genomics Consortium, Department of Medical Biochemistry and Biophysics, Karolinska Institute, Stockholm, Sweden.
Organizational Affiliation: