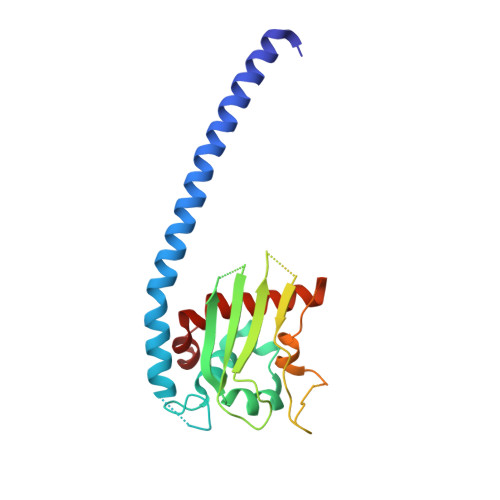
Iodide-SAD, SIR and SIRAS phasing for structure solution of a nucleosome assembly protein.
Yogavel, M., Gill, J., Sharma, A.(2009) Acta Crystallogr D Biol Crystallogr 65: 618-622
- PubMed: 19465776 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909013171
- Primary Citation Related Structures:
3GYV, 3GYW - PubMed Abstract:
The crystal structure of Plasmodium falciparum nucleosome assembly protein (PfNapL) was determined by iodide-SAD/SIRAS phasing methods using iodide-SAD data to 3.0 A resolution and native data to 2.4 A resolution. Halide-derivatized PfNapL crystals were obtained using the quick cryo-soaking method in which the native crystals were soaked in a cryosolution consisting of 500 mM NaI for a short period of 30-60 s and data were collected at an in-house X-ray source using Cu Kalpha radiation. Despite a low anomalous signal-to-noise ratio of <1.2 in the >3.5 A resolution bin, the data were sufficient to determine the structure by SAD/SIR/SIRAS methods using the soaked iodides. Previously, structure solution had failed with both molecular-replacement and selenomethionine-derivatization techniques owing to reasons that are detailed in this work. The phasing at low resolution with three iodides per monomer with high temperature factors was successful using any of the SAD, SIR or SIRAS methods.
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), New Delhi 110 067, India.
Organizational Affiliation: