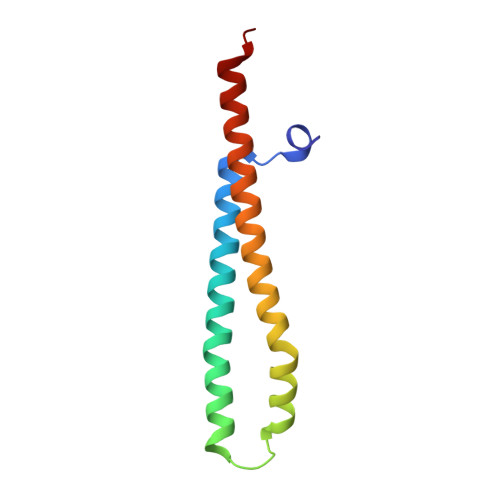
WXG100 protein superfamily consists of three subfamilies and exhibits an alpha-helical C-terminal conserved residue pattern.
Poulsen, C., Panjikar, S., Holton, S.J., Wilmanns, M., Song, Y.H.(2014) PLoS One 9: e89313-e89313
- PubMed: 24586681
- DOI: https://doi.org/10.1371/journal.pone.0089313
- Primary Citation Related Structures:
3FAV, 3GVM, 3GWK - PubMed Abstract:
Members of the WXG100 protein superfamily form homo- or heterodimeric complexes. The most studied proteins among them are the secreted T-cell antigens CFP-10 (10 kDa culture filtrate protein, EsxB) and ESAT-6 (6 kDa early secreted antigen target, EsxA) from Mycobacterium tuberculosis. They are encoded on an operon within a gene cluster, named as ESX-1, that encodes for the Type VII secretion system (T7SS). WXG100 proteins are secreted in a full-length form and it is known that they adopt a four-helix bundle structure. In the current work we discuss the evolutionary relationship between the homo- and heterodimeric WXG100 proteins, the basis of the oligomeric state and the key structural features of the conserved sequence pattern of WXG100 proteins. We performed an iterative bioinformatics analysis of the WXG100 protein superfamily and correlated this with the atomic structures of the representative WXG100 proteins. We find, firstly, that the WXG100 protein superfamily consists of three subfamilies: CFP-10-, ESAT-6- and sagEsxA-like proteins (EsxA proteins similar to that of Streptococcus agalactiae). Secondly, that the heterodimeric complexes probably evolved from a homodimeric precursor. Thirdly, that the genes of hetero-dimeric WXG100 proteins are always encoded in bi-cistronic operons and finally, by combining the sequence alignments with the X-ray data we identify a conserved C-terminal sequence pattern. The side chains of these conserved residues decorate the same side of the C-terminal α-helix and therefore form a distinct surface. Our results lead to a putatively extended T7SS secretion signal which combines two reported T7SS recognition characteristics: Firstly that the T7SS secretion signal is localized at the C-terminus of T7SS substrates and secondly that the conserved residues YxxxD/E are essential for T7SS activity. Furthermore, we propose that the specific α-helical surface formed by the conserved sequence pattern including YxxxD/E motif is a key component of T7SS-substrate recognition.
- EMBL-Hamburg, Hamburg, Germany.
Organizational Affiliation: