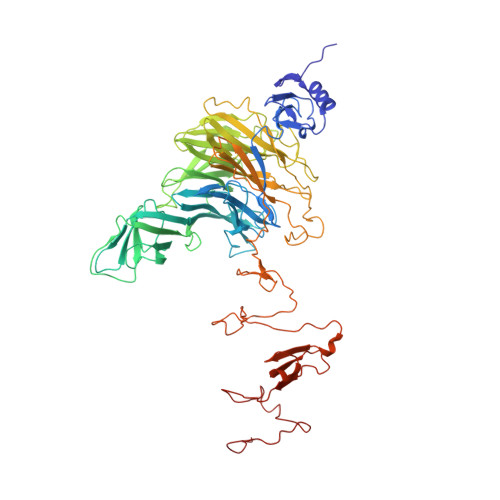
Structural basis for the recognition and cleavage of polysialic acid by the bacteriophage K1F tailspike protein EndoNF.
Schulz, E.C., Schwarzer, D., Frank, M., Stummeyer, K., Muhlenhoff, M., Dickmanns, A., Gerardy-Schahn, R., Ficner, R.(2010) J Mol Biology 397: 341-351
- PubMed: 20096705 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.01.028
- Primary Citation Related Structures:
3GVJ, 3GVK, 3GVL - PubMed Abstract:
An alpha-2,8-linked polysialic acid (polySia) capsule confers immune tolerance to neuroinvasive, pathogenic prokaryotes such as Escherichia coli K1 and Neisseria meningitidis and supports host infection by means of molecular mimicry. Bacteriophages of the K1 family, infecting E. coli K1, specifically recognize and degrade this polySia capsule utilizing tailspike endosialidases. While the crystal structure for the catalytic domain of the endosialidase of bacteriophage K1F (endoNF) has been solved, there is yet no structural information on the mode of polySia binding and cleavage available. The crystal structure of activity deficient active-site mutants of the homotrimeric endoNF cocrystallized with oligomeric sialic acid identified three independent polySia binding sites in each endoNF monomer. The bound oligomeric sialic acid displays distinct conformations at each site. In the active site, a Sia(3) molecule is bound in an extended conformation representing the enzyme-product complex. Structural and biochemical data supported by molecular modeling enable to propose a reaction mechanism for polySia cleavage by endoNF.
- Abteilung für Molekulare Strukturbiologie, Institut für Mikrobiologie und Genetik, Georg-August-Universität Göttingen, Justus-von-Liebig-Weg 11, 37077 Göttingen, Germany.
Organizational Affiliation: