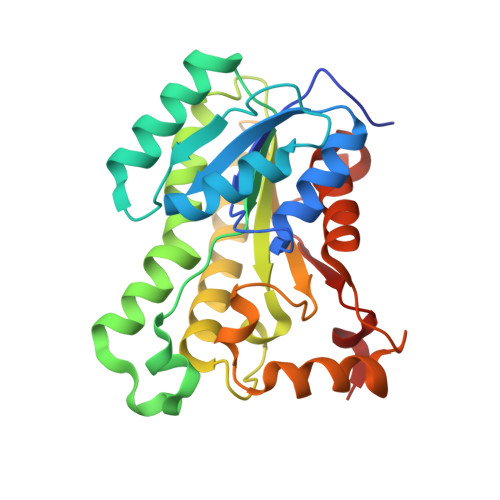
Structural insights into Staphylococcus aureus enoyl-ACP reductase (FabI), in complex with NADP and triclosan.
Priyadarshi, A., Kim, E.E., Hwang, K.Y.(2010) Proteins 78: 480-486
- PubMed: 19768684 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22581
- Primary Citation Related Structures:
3GNS, 3GNT, 3GR6 - Division of Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 136-701, South Korea.
Organizational Affiliation: