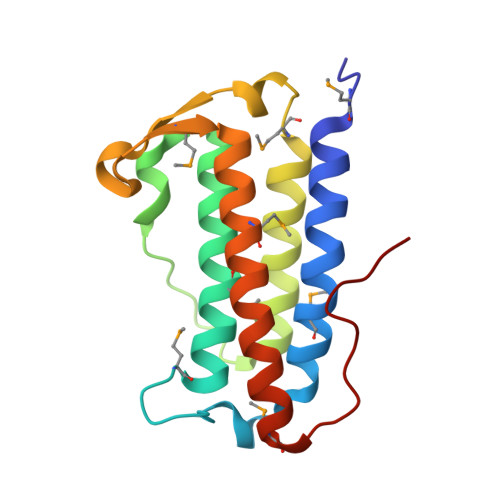
The structure of DinB from Geobacillus stearothermophilus: a representative of a unique four-helix-bundle superfamily.
Cooper, D.R., Grelewska, K., Kim, C.Y., Joachimiak, A., Derewenda, Z.S.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 219-224
- PubMed: 20208147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109053913
- Primary Citation Related Structures:
3GOR - PubMed Abstract:
The crystal structure of the dinB gene product from Geobacillus stearothermophilus (GsDinB) is reported at 2.5 A resolution. The dinB gene is one of the DNA-damage-induced genes and the corresponding protein, DinB, is the founding member of a Pfam family with no known function. The protein contains a four-helix up-down-down-up bundle that has previously been described in the literature in three disparate proteins: the enzyme MDMPI (mycothiol-dependent maleylpyruvate isomerase), YfiT and TTHA0303, a member of a small DUF (domain of unknown function). However, a search of the DALI structural database revealed similarities to a further 11 new unpublished structures contributed by structural genomics centers. The sequences of these proteins are quite divergent and represent several Pfam families, yet their structures are quite similar and most (but not all) seem to have the ability to coordinate a metal ion using a conserved histidine-triad motif. The structural similarities of these diverse proteins suggest that a new Pfam clan encompassing the families that share this fold should be created. The proteins that share this fold exhibit four different quaternary structures: monomeric and three different dimeric forms.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, Virginia 22908, USA.
Organizational Affiliation: