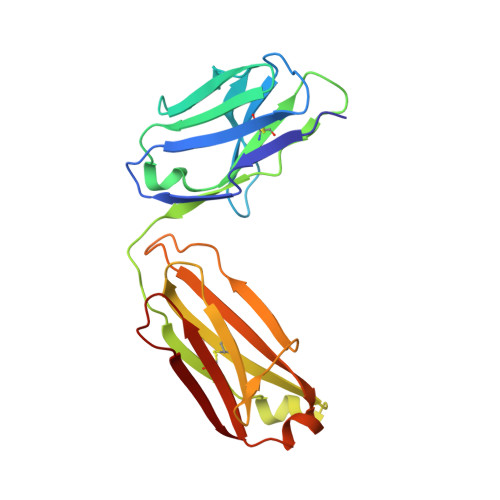
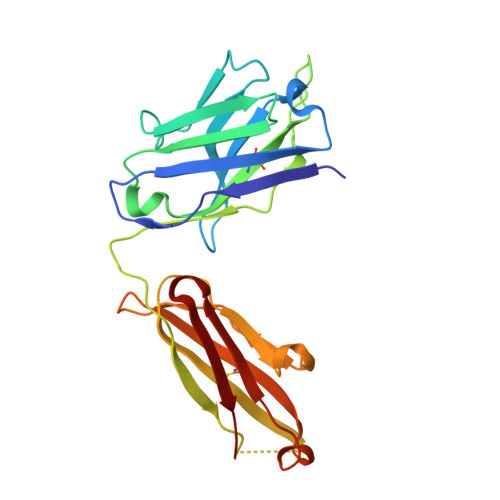
Structure of the Fab fragment of therapeutic antibody Ofatumumab provides insights into the recognition mechanism with CD20
Du, J., Yang, H., Guo, Y., Ding, J.(2009) Mol Immunol 46: 2419-2423
- PubMed: 19427037
- DOI: https://doi.org/10.1016/j.molimm.2009.04.009
- Primary Citation Related Structures:
3GIZ - PubMed Abstract:
CD20 is an important drug target for B-cell depletion therapy against certain B-cell lymphomas and autoimmune diseases. The success of anti-CD20 antibody drugs such as Rituximab, Ibritumomab, and Tositumomab has promoted the development of new generation of anti-CD20 antibodies for therapeutic applications. Ofatumumab is a fully human anti-CD20 antibody that is currently in phase III clinical trial for several types of malignancies and autoimmune diseases and is one of the most promising anti-CD20 drugs. Here we report the crystal structure of the Fab fragment of Ofatumumab at 2.2A resolution. The antigen combining site is composed of a large, deep pocket formed by six CDR loops. The pocket has a hydrophobic periphery and a positively charged bottom. Structure analysis and comparison with other antibodies suggest that the hydrophobic periphery might interact with the epitope on CD20 that is enriched with hydrophobic residues and very close to cell membrane, and the positively charged bottom might interact with Glu(150) of CD20 which is the only negatively charged residue within the epitope. These results provide some insights into the recognition of Ofatumumab with CD20 and explain how the antibody can recognize an epitope so close to the cell membrane.
- State Key Laboratory of Molecular Biology and Research Center for Structural Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.
Organizational Affiliation: