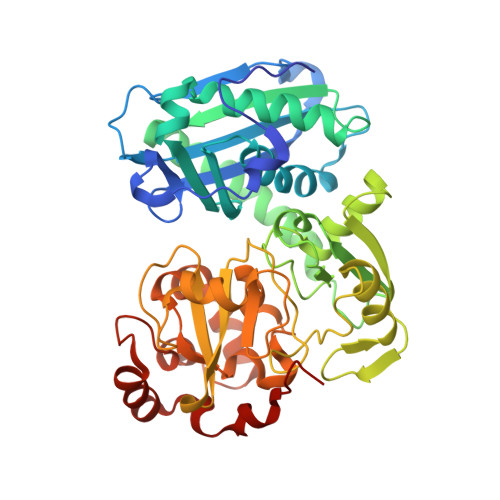
Novel domain arrangement in the crystal structure of a truncated acetyl-CoA synthase from Moorella thermoacetica
Volbeda, A., Darnault, C., Tan, X., Lindahl, P.A., Fontecilla-Camps, J.C.(2009) Biochemistry 48: 7916-7926
- PubMed: 19650626
- DOI: https://doi.org/10.1021/bi9003952
- Primary Citation Related Structures:
3GIT - PubMed Abstract:
Ni-dependent acetyl-CoA synthase (ACS) and CO dehydrogenase (CODH) constitute the central enzyme complex of the Wood-Ljungdahl pathway of acetyl-CoA formation. The crystal structure of a recombinant bacterial ACS lacking the N-terminal domain that interacts with CODH shows a large reorganization of the remaining two globular domains, producing a narrow cleft of suitable size, shape, and nature to bind CoA. Sequence comparisons with homologous archaeal enzymes that naturally lack the N-terminal domain show that many amino acids lining this cleft are conserved. Besides the typical [4Fe-4S] center, the A-cluster contains only one proximal metal ion that, according to anomalous scattering data, is most likely Cu or Zn. Incorporation of a functional Ni(2)Fe(4)S(4) A-cluster would require only minor structural rearrangements. Using available structures, a plausible model of the interaction between CODH and the smaller ACS in archaeal multienzyme complexes is presented, along with a discussion of evolutionary relationships of the archaeal and bacterial enzymes.
- Laboratoire de Cristallographie et Cristallogenèse des Protéines, Institut de Biologie Structurale Jean-Pierre Ebel, CEA, CNRS, Université Joseph Fourier, 41 Rue Jules Horowitz, F-38027 Grenoble, France. anne.volbeda@ibs.fr
Organizational Affiliation: