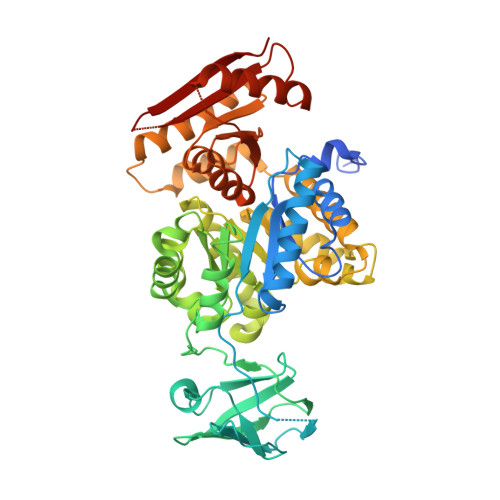
The crystal structure of Toxoplasma gondii pyruvate kinase 1.
Bakszt, R., Wernimont, A., Allali-Hassani, A., Mok, M.W., Hills, T., Hui, R., Pizarro, J.C.(2010) PLoS One 5: e12736-e12736
- PubMed: 20856875
- DOI: https://doi.org/10.1371/journal.pone.0012736
- Primary Citation Related Structures:
3EOE, 3GG8 - PubMed Abstract:
Pyruvate kinase (PK), which catalyzes the final step in glycolysis converting phosphoenolpyruvate to pyruvate, is a central metabolic regulator in most organisms. Consequently PK represents an attractive therapeutic target in cancer and human pathogens, like Apicomplexans. The phylum Aplicomplexa, a group of exclusively parasitic organisms, includes the genera Plasmodium, Cryptosporidium and Toxoplasma, the etiological agents of malaria, cryptosporidiosis and toxoplasmosis respectively. Toxoplasma gondii infection causes a mild illness and is a very common infection affecting nearly one third of the world's population. We have determined the crystal structure of the PK1 enzyme from T. gondii, with the B domain in the open and closed conformations. We have also characterized its enzymatic activity and confirmed glucose-6-phosphate as its allosteric activator. This is the first description of a PK enzyme in a closed inactive conformation without any bound substrate. Comparison of the two tetrameric TgPK1 structures indicates a reorientation of the monomers with a concomitant change in the buried surface among adjacent monomers. The change in the buried surface was associated with significant B domain movements in one of the interacting monomers. We hypothesize that a loop in the interface between the A and B domains plays an important role linking the position of the B domain to the buried surface among monomers through two α-helices. The proposed model links the catalytic cycle of the enzyme with its domain movements and highlights the contribution of the interface between adjacent subunits. In addition, an unusual ordered conformation was observed in one of the allosteric binding domains and it is related to a specific apicomplexan insertion. The sequence and structural particularity would explain the atypical activation by a mono-phosphorylated sugar. The sum of peculiarities raises this enzyme as an emerging target for drug discovery.
- The Structural Genomics Consortium, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: