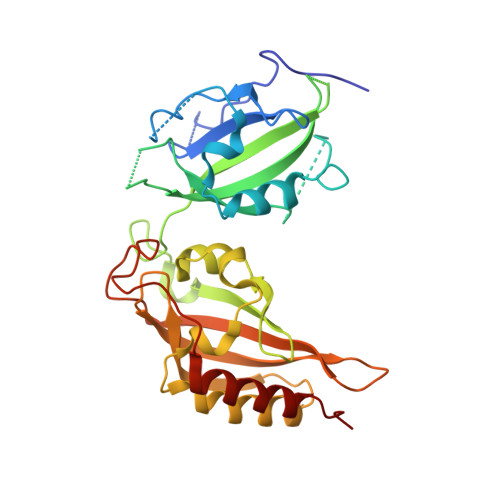
Structural and functional analyses of PAS domain interactions of the clock proteins Drosophila PERIOD and mouse PERIOD2.
Hennig, S., Strauss, H.M., Vanselow, K., Yildiz, O., Schulze, S., Arens, J., Kramer, A., Wolf, E.(2009) PLoS Biol 7: e94-e94
- PubMed: 19402751 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1000094
- Primary Citation Related Structures:
3GDI, 3GEC - PubMed Abstract:
PERIOD proteins are central components of the Drosophila and mammalian circadian clocks. The crystal structure of a Drosophila PERIOD (dPER) fragment comprising two PER-ARNT-SIM (PAS) domains (PAS-A and PAS-B) and two additional C-terminal alpha-helices (alphaE and alphaF) has revealed a homodimer mediated by intermolecular interactions of PAS-A with tryptophane 482 in PAS-B and helix alphaF. Here we present the crystal structure of a monomeric PAS domain fragment of dPER lacking the alphaF helix. Moreover, we have solved the crystal structure of a PAS domain fragment of the mouse PERIOD homologue mPER2. The mPER2 structure shows a different dimer interface than dPER, which is stabilized by interactions of the PAS-B beta-sheet surface including tryptophane 419 (equivalent to Trp482dPER). We have validated and quantitatively analysed the homodimer interactions of dPER and mPER2 by site-directed mutagenesis using analytical gel filtration, analytical ultracentrifugation, and co-immunoprecipitation experiments. Furthermore we show, by yeast-two-hybrid experiments, that the PAS-B beta-sheet surface of dPER mediates interactions with TIMELESS (dTIM). Our study reveals quantitative and qualitative differences between the homodimeric PAS domain interactions of dPER and its mammalian homologue mPER2. In addition, we identify the PAS-B beta-sheet surface as a versatile interaction site mediating mPER2 homodimerization in the mammalian system and dPER-dTIM heterodimer formation in the Drosophila system.
- Max Planck Institute of Molecular Physiology, Department of Structural Biology, Dortmund, Germany.
Organizational Affiliation: