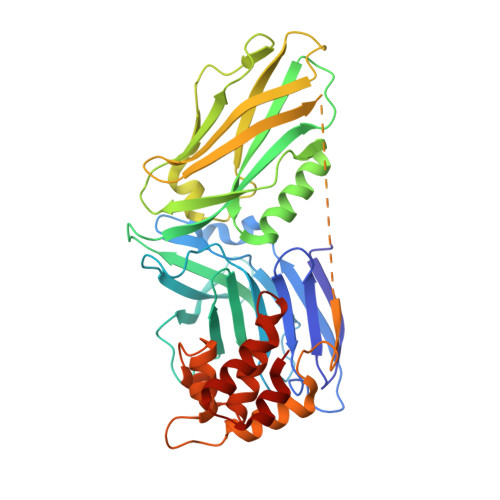
Autoinhibition of UNC5b revealed by the cytoplasmic domain structure of the receptor
Wang, R., Wei, Z., Jin, H., Wu, H., Yu, C., Wen, W., Chan, L.-N., Wen, Z., Zhang, M.(2009) Mol Cell 33: 692-703
- PubMed: 19328064 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2009.02.016
- Primary Citation Related Structures:
3G5B - PubMed Abstract:
The cytoplasmic domains of UNC5 are responsible for its netrin-mediated signaling events in axonal migrations, blood vessel patterning, and apoptosis, although the molecular mechanisms governing these processes are unknown. To provide a foundation for the elucidation of the UNC5-mediated signaling mechanism, we determined the crystal structure of the cytoplasmic portion of UNC5b. We found that it contains three distinctly folded domains, namely ZU5, UPA, and death domain (DD). These three domains form a structural supramodule, with ZU5 binding to both UPA and DD, thereby locking the ZU5-UPA-DD supramodule in a closed conformation and suppressing its biological activities. Release of the closed conformation of the ZU5-UPA-DD supramodule leads to the activation of the receptor in the promotion of apoptosis and blood vessel patterning. Finally, we provide evidence showing that the supramodular nature of UNC5 ZU5-UPA-DD is likely to be shared by the ankyrin and PIDD families of scaffold proteins.
- Department of Biochemistry, Molecular Neuroscience Center, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong.
Organizational Affiliation: