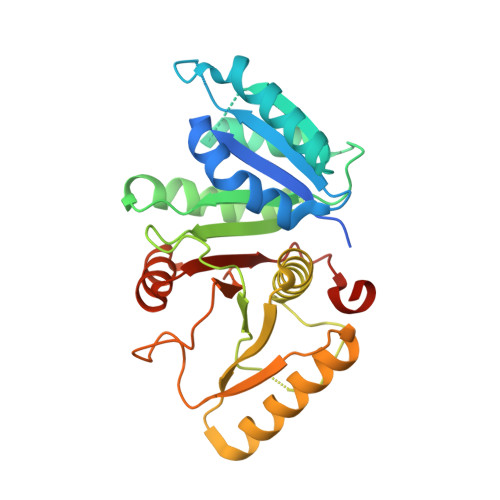
X-ray Structure of the C-Terminal Domain of a Prokaryotic Cation-Chloride Cotransporter
Warmuth, S., Zimmermann, I., Dutzler, R.(2009) Structure 17: 538-546
- PubMed: 19368887 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.02.009
- Primary Citation Related Structures:
3G40 - PubMed Abstract:
The cation-chloride cotransporters (CCCs) mediate the electroneutral transport of chloride in dependence of sodium and potassium. The proteins share a conserved structural scaffold that consists of a transmembrane transport domain followed by a cytoplasmic regulatory domain. We have determined the X-ray structure of the C-terminal domain of the archaea Methanosarcina acetivorans. The structure shows a novel fold of a regulatory domain that is distantly related to universal stress proteins. The protein forms dimers in solution, which is consistent with the proposed dimeric organization of eukaryotic CCC transporters. The dimer interface observed in different crystal forms is unusual because the buried area is relatively small and hydrophilic. By using a biochemical approach we show that this interaction is preserved in solution and in the context of the full-length transporter. Our studies reveal structural insight into the CCC family and establish the oligomeric organization of this important class of transport proteins.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland.
Organizational Affiliation: