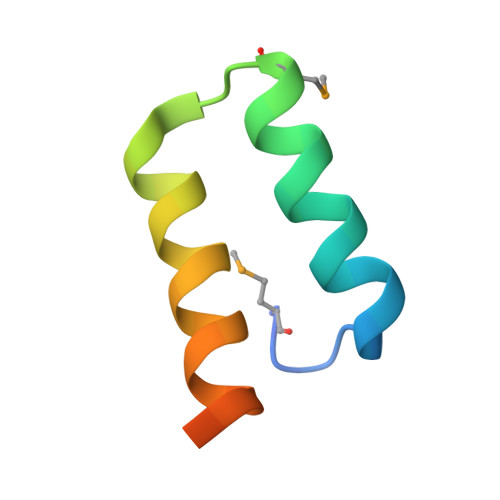
Structure of the sporulation histidine kinase inhibitor Sda from Bacillus subtilis and insights into its solution state
Jacques, D.A., Streamer, M., Rowland, S.L., King, G.F., Guss, J.M., Trewhella, J., Langley, D.B.(2009) Acta Crystallogr D Biol Crystallogr 65: 574-581
- PubMed: 19465772
- DOI: https://doi.org/10.1107/S090744490901169X
- Primary Citation Related Structures:
3FYR - PubMed Abstract:
The crystal structure of the DNA-damage checkpoint inhibitor of sporulation, Sda, from Bacillus subtilis, has been solved by the MAD technique using selenomethionine-substituted protein. The structure closely resembles that previously solved by NMR, as well as the structure of a homologue from Geobacillus stearothermophilus solved in complex with the histidine kinase KinB. The structure contains three molecules in the asymmetric unit. The unusual trimeric arrangement, which lacks simple internal symmetry, appears to be preserved in solution based on an essentially ideal fit to previously acquired scattering data for Sda in solution. This interpretation contradicts previous findings that Sda was monomeric or dimeric in solution. This study demonstrates the difficulties that can be associated with the characterization of small proteins and the value of combining multiple biophysical techniques. It also emphasizes the importance of understanding the physical principles behind these techniques and therefore their limitations.
- School of Molecular and Microbial Biosciences, University of Sydney, Australia.
Organizational Affiliation: