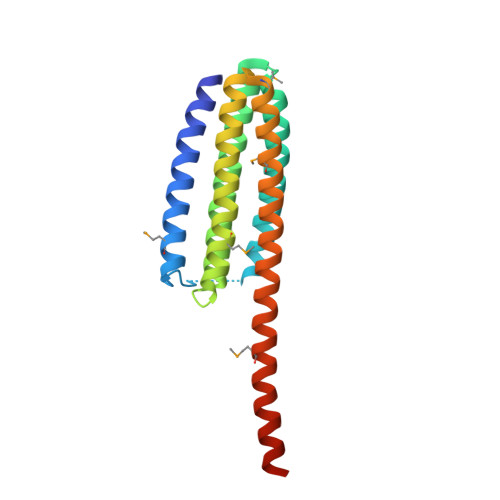
Crystal structure of the talin integrin binding domain 2.
Cheung, T.Y., Fairchild, M.J., Zarivach, R., Tanentzapf, G., Van Petegem, F.(2009) J Mol Biology 387: 787-793
- PubMed: 19340939 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.01.053
- Primary Citation Related Structures:
3FYQ - PubMed Abstract:
Integrins are transmembrane receptors that mediate cell adhesion to the extracellular matrix and play essential roles in tissue development and maintenance. The cytoplasmic segment of integrin associates with talin, a large intracellular protein that links integrin to the actin cytoskeleton. Binding of talin via an integrin binding segment (IBS1) results in large conformational changes in the extracellular portion of integrin, which modulates the affinity of integrins for their extracellular matrix ligands. However, integrin binding also requires a second segment of talin (IBS2). Despite detailed descriptions of the integrin-IBS1 binding, the molecular determinants that drive the integrin-IBS2 association are poorly understood. Here, we describe the crystal structure of the talin IBS2 domain, which forms a five-helix bundle. The large structural homology with a vinculin binding domain hints at an ancient gene duplication and suggests that helix 4 may bind to vinculin if the bundle is unfolded. Mapping previous mutations on the surface highlights a likely binding interface for integrin.
- Department of Cellular and Physiological Sciences, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: