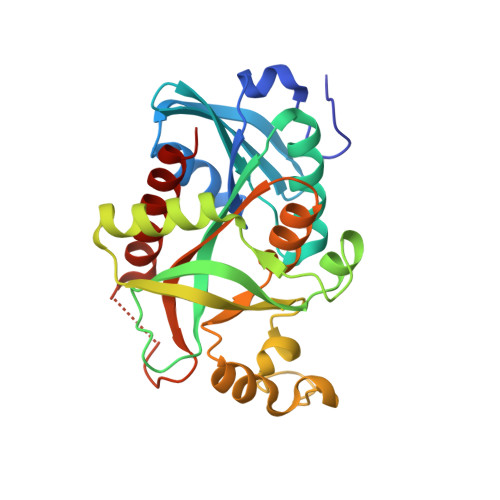
The X-ray structure of Salmonella typhimurium uridine nucleoside phosphorylase complexed with 2,2'-anhydrouridine, phosphate and potassium ions at 1.86 A resolution.
Lashkov, A.A., Zhukhlistova, N.E., Gabdoulkhakov, A.H., Shtil, A.A., Efremov, R.G., Betzel, C., Mikhailov, A.M.(2010) Acta Crystallogr D Biol Crystallogr 66: 51-60
- PubMed: 20057049 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909044175
- Primary Citation Related Structures:
3FWP - PubMed Abstract:
Uridine nucleoside phosphorylase is an important drug target for the development of anti-infective and antitumour agents. The X-ray crystal structure of Salmonella typhimurium uridine nucleoside phosphorylase (StUPh) complexed with its inhibitor 2,2'-anhydrouridine, phosphate and potassium ions has been solved and refined at 1.86 A resolution (R(cryst) = 17.6%, R(free) = 20.6%). The complex of human uridine phosphorylase I (HUPhI) with 2,2'-anhydrouridine was modelled using a computational approach. The model allowed the identification of atomic groups in 2,2'-anhydrouridine that might improve the interaction of future inhibitors with StUPh and HUPhI.
- A. V. Shubnikov Institute of Crystallography, Russian Academy of Sciences, 59 Leninsky Prospect, 119333 Moscow, Russia.
Organizational Affiliation: