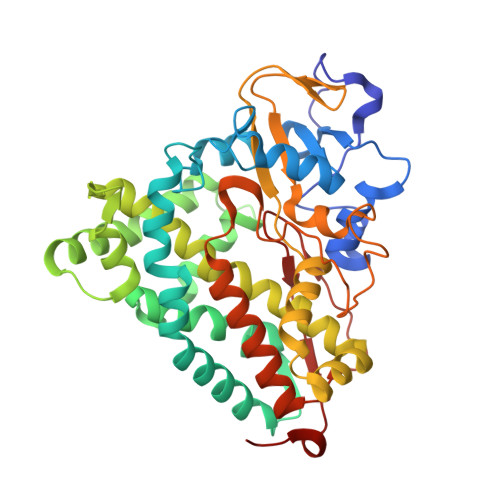
Probing the role of the proximal heme ligand in cytochrome P450cam by recombinant incorporation of selenocysteine.
Aldag, C., Gromov, I.A., Garcia-Rubio, I., von Koenig, K., Schlichting, I., Jaun, B., Hilvert, D.(2009) Proc Natl Acad Sci U S A 106: 5481-5486
- PubMed: 19293375 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0810503106
- Primary Citation Related Structures:
3FWF, 3FWG, 3FWI, 3FWJ - PubMed Abstract:
The unique monooxygenase activity of cytochrome P450cam has been attributed to coordination of a cysteine thiolate to the heme cofactor. To investigate this interaction, we replaced cysteine with the more electron-donating selenocysteine. Good yields of the selenoenzyme were obtained by bacterial expression of an engineered gene containing the requisite UGA codon for selenocysteine and a simplified yet functional selenocysteine insertion sequence (SECIS). The sulfur-to-selenium substitution subtly modulates the structural, electronic, and catalytic properties of the enzyme. Catalytic activity decreases only 2-fold, whereas substrate oxidation becomes partially uncoupled from electron transfer, implying a more complex role for the axial ligand than generally assumed.
- Laboratory of Organic Chemistry, ETH Zurich, CH-8093 Zurich, Switzerland.
Organizational Affiliation: