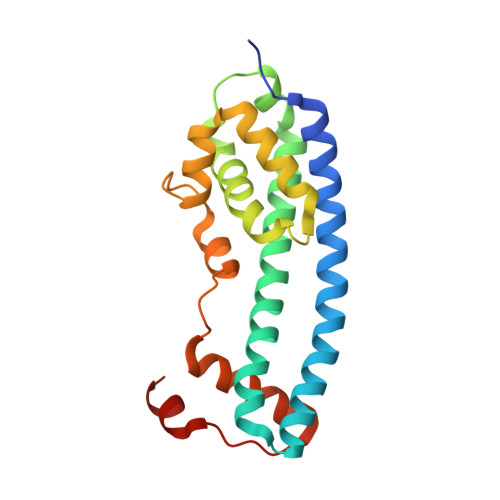
Structural basis for ESCRT-III protein autoinhibition.
Bajorek, M., Schubert, H.L., McCullough, J., Langelier, C., Eckert, D.M., Stubblefield, W.M., Uter, N.T., Myszka, D.G., Hill, C.P., Sundquist, W.I.(2009) Nat Struct Mol Biol 16: 754-762
- PubMed: 19525971
- DOI: https://doi.org/10.1038/nsmb.1621
- Primary Citation of Related Structures:
3FRR, 3FRS, 3FRT, 3FRV - PubMed Abstract:
Endosomal sorting complexes required for transport-III (ESCRT-III) subunits cycle between two states: soluble monomers and higher-order assemblies that bind and remodel membranes during endosomal vesicle formation, midbody abscission and enveloped virus budding. Here we show that the N-terminal core domains of increased sodium tolerance-1 (IST1) and charged multivesicular body protein-3 (CHMP3) form equivalent four-helix bundles, revealing that IST1 is a previously unrecognized ESCRT-III family member. IST1 and its ESCRT-III binding partner, CHMP1B, both form higher-order helical structures in vitro, and IST1-CHMP1 interactions are required for abscission. The IST1 and CHMP3 structures also reveal that equivalent downstream alpha5 helices can fold back against the core domains. Mutations within the CHMP3 core-alpha5 interface stimulate the protein's in vitro assembly and HIV-inhibition activities, indicating that dissociation of the autoinhibitory alpha5 helix from the core activates ESCRT-III proteins for assembly at membranes.
- Department of Biochemistry, University of Utah, Salt Lake City, Utah, USA.
Organizational Affiliation: