Quinalizarin as a potent, selective and cell-permeable inhibitor of protein kinase CK2
Cozza, G., Mazzorana, M., Papinutto, E., Bain, J., Elliott, M., di Maira, G., Gianoncelli, A., Pagano, M.A., Sarno, S., Ruzzene, M., Battistutta, R., Meggio, F., Moro, S., Zagotto, G., Pinna, L.A.(2009) Biochem J 421: 387-395
- PubMed: 19432557 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20090069
- Primary Citation Related Structures:
3FL5 - PubMed Abstract:
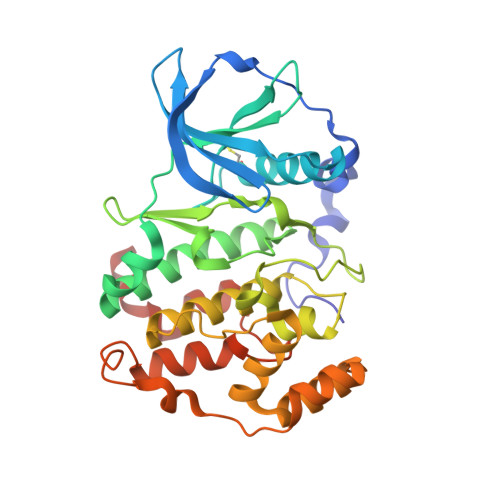
Emodin (1,3,8-trihydroxy-6-methyl-anthraquinone) is a moderately potent and poorly selective inhibitor of protein kinase CK2, one of the most pleiotropic serine/threonine protein kinases, implicated in neoplasia and in other global diseases. By virtual screening of the MMS (Molecular Modeling Section) database, we have now identified quinalizarin (1,2,5,8-tetrahydroxyanthraquinone) as an inhibitor of CK2 that is more potent and selective than emodin. CK2 inhibition by quinalizarin is competitive with respect to ATP, with a Ki value of approx. 50 nM. Tested at 1 microM concentration on a panel of 75 protein kinases, quinalizarin drastically inhibits only CK2, with a promiscuity score (11.1), which is the lowest ever reported so far for a CK2 inhibitor. Especially remarkable is the ability of quinalizarin to discriminate between CK2 and a number of kinases, notably DYRK1a (dual-specificity tyrosine-phosphorylated and -regulated kinase), PIM (provirus integration site for Moloney murine leukaemia virus) 1, 2 and 3, HIPK2 (homeodomain-interacting protein kinase-2), MNK1 [MAPK (mitogen-activated protein kinase)-interacting kinase 1], ERK8 (extracellular-signal-regulated kinase 8) and PKD1 (protein kinase D 1), which conversely tend to be inhibited as drastically as CK2 by commercially available CK2 inhibitors. The determination of the crystal structure of a complex between quinalizarin and CK2alpha subunit highlights the relevance of polar interactions in stabilizing the binding, an unusual characteristic for a CK2 inhibitor, and disclose other structural features which may account for the narrow selectivity of this compound. Tested on Jurkat cells, quinalizarin proved able to inhibit endogenous CK2 and to induce apoptosis more efficiently than the commonly used CK2 inhibitors TBB (4,5,6,7-tetrabromo-1H-benzotriazole) and DMAT (2-dimethylamino-4,5,6,7-tetrabromo-1H-benzimidazole).
- Department of Biological Chemistry and CNR Institute of Neurosciences, University of Padova, viale G. Colombo 3, 35121 Padova, Italy.
Organizational Affiliation: