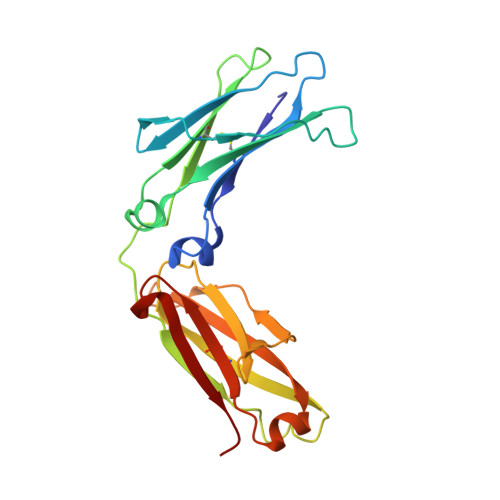
Structural characterization of a human Fc fragment engineered for extended serum half-life.
Oganesyan, V., Damschroder, M.M., Woods, R.M., Cook, K.E., Wu, H., Dall'acqua, W.F.(2009) Mol Immunol 46: 1750-1755
- PubMed: 19250681 Search on PubMed
- DOI: https://doi.org/10.1016/j.molimm.2009.01.026
- Primary Citation Related Structures:
3FJT - PubMed Abstract:
The first three-dimensional structure of a human Fc fragment genetically engineered for improved pharmacokinetics properties is reported. When introduced into the C(H)2 domain of human immunoglobulin G (IgG) molecules, the triple mutation M252Y/S254T/T256E ('YTE') causes an about 10-fold increase in their binding to the human neonatal Fc receptor (FcRn). This translates into an almost 4-fold increase in the serum half-life of YTE-containing human IgGs in cynomolgus monkeys. A recombinantly produced human Fc/YTE fragment was crystallized and its structure solved at a resolution of 2.5A using molecular replacement. This revealed that Fc/YTE three-dimensional structure is very similar to that of other human Fc fragments in the experimentally visible region spanning residues 236-444. We propose that the enhanced interaction between Fc/YTE and human FcRn is likely mediated by local effects at the substitutions sites. Molecular modeling suggested that potential favorable hydrogen bonds along with an increase in the surface of contact between the two partners may account in part for the corresponding increase in affinity.
- Department of Antibody Discovery and Protein Engineering, MedImmune, One MedImmune Way, Gaithersburg, MD 20878, USA.
Organizational Affiliation: