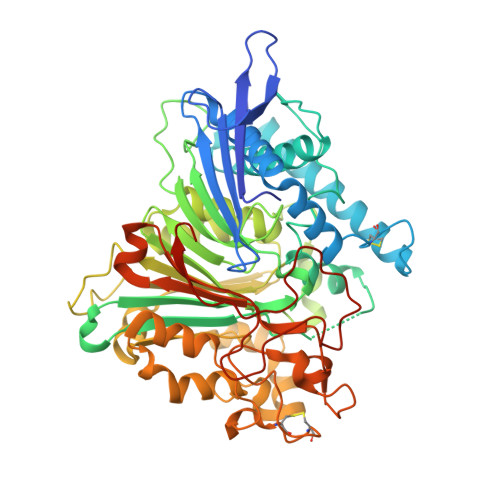
Initial insight into the function of the lysosomal 66.3 kDa protein from mouse by means of X-ray crystallography
Lakomek, K., Dickmanns, A., Kettwig, M., Urlaub, H., Ficner, R., Luebke, T.(2009) BMC Struct Biol 9: 56-56
- PubMed: 19706171 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-9-56
- Primary Citation Related Structures:
3FGR, 3FGT, 3FGW - PubMed Abstract:
The lysosomal 66.3 kDa protein from mouse is a soluble, mannose 6-phosphate containing protein of so far unknown function. It is synthesized as a glycosylated 75 kDa precursor that undergoes limited proteolysis leading to a 28 kDa N- and a 40 kDa C-terminal fragment. In order to gain insight into the function and the post-translational maturation process of the glycosylated 66.3 kDa protein, three crystal structures were determined that represent different maturation states. These structures demonstrate that the 28 kDa and 40 kDa fragment which have been derived by a proteolytic cleavage remain associated. Mass spectrometric analysis confirmed the subsequent trimming of the C-terminus of the 28 kDa fragment making a large pocket accessible, at the bottom of which the putative active site is located. The crystal structures reveal a significant similarity of the 66.3 kDa protein to several bacterial hydrolases. The core alphabetabetaalpha sandwich fold and a cysteine residue at the N-terminus of the 40 kDa fragment (C249) classify the 66.3 kDa protein as a member of the structurally defined N-terminal nucleophile (Ntn) hydrolase superfamily. Due to the close resemblance of the 66.3 kDa protein to members of the Ntn hydrolase superfamily a hydrolytic activity on substrates containing a non-peptide amide bond seems reasonable. The structural homology which comprises both the overall fold and essential active site residues also implies an autocatalytic maturation process of the lysosomal 66.3 kDa protein. Upon the proteolytic cleavage between S248 and C249, a deep pocket becomes solvent accessible, which harbors the putative active site of the 66.3 kDa protein.
- Department of Molecular Structural Biology, Institute of Microbiology and Genetics, GZMB, Georg-August University Goettingen, Justus-von-Liebig-Weg 11, D-37077 Goettingen, Germany. klakome@gwdg.de
Organizational Affiliation: