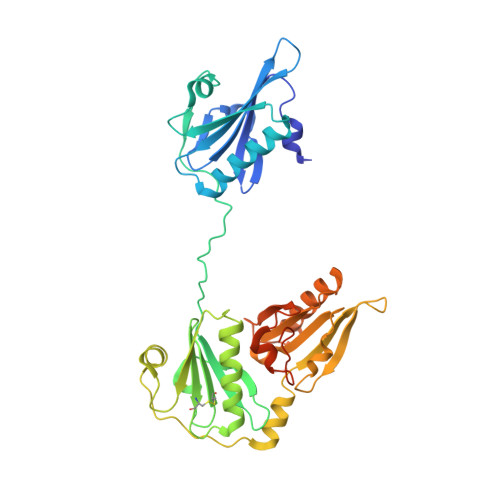
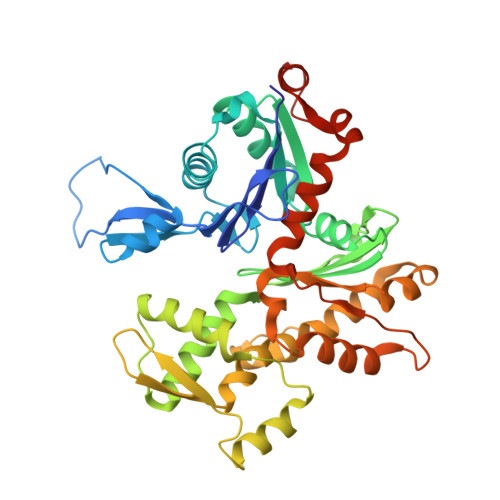
Ca2+ binding by domain 2 plays a critical role in the activation and stabilization of gelsolin.
Nag, S., Ma, Q., Wang, H., Chumnarnsilpa, S., Lee, W.L., Larsson, M., Kannan, B., Hernandez-Valladares, M., Burtnick, L.D., Robinson, R.C.(2009) Proc Natl Acad Sci U S A 106: 13713-13718
- PubMed: 19666512 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0812374106
- Primary Citation Related Structures:
3FFK, 3FFN - PubMed Abstract:
Gelsolin consists of six homologous domains (G1-G6), each containing a conserved Ca-binding site. Occupation of a subset of these sites enables gelsolin to sever and cap actin filaments in a Ca-dependent manner. Here, we present the structures of Ca-free human gelsolin and of Ca-bound human G1-G3 in a complex with actin. These structures closely resemble those determined previously for equine gelsolin. However, the G2 Ca-binding site is occupied in the human G1-G3/actin structure, whereas it is vacant in the equine version. In-depth comparison of the Ca-free and Ca-activated, actin-bound human gelsolin structures suggests G2 and G6 to be cooperative in binding Ca(2+) and responsible for opening the G2-G6 latch to expose the F-actin-binding site on G2. Mutational analysis of the G2 and G6 Ca-binding sites demonstrates their interdependence in maintaining the compact structure in the absence of calcium. Examination of Ca binding by G2 in human G1-G3/actin reveals that the Ca(2+) locks the G2-G3 interface. Thermal denaturation studies of G2-G3 indicate that Ca binding stabilizes this fragment, driving it into the active conformation. The G2 Ca-binding site is mutated in gelsolin from familial amyloidosis (Finnish-type) patients. This disease initially proceeds through protease cleavage of G2, ultimately to produce a fragment that forms amyloid fibrils. The data presented here support a mechanism whereby the loss of Ca binding by G2 prolongs the lifetime of partially activated, intermediate conformations in which the protease cleavage site is exposed.
- Institute of Molecular and Cell Biology, ASTAR, 61 Biopolis Drive, Proteos, Singapore 138673.
Organizational Affiliation: