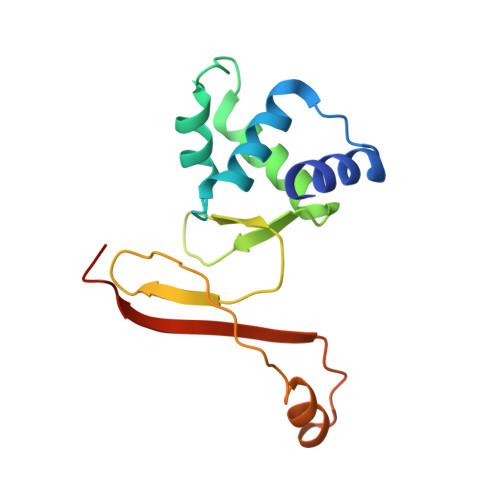
Structural characterization of the active form of PerR: insights into the metal-induced activation of PerR and Fur proteins for DNA binding
Jacquamet, L., Traore, D.A.K., Ferrer, J.-L., Proux, O., Testemale, D., Hazemann, J.-L., Nazarenko, E., El Ghazouani, A., Caux-Thang, C., Duarte, V., Latour, J.-M.(2009) Mol Microbiol 73: 20-31
- PubMed: 19508285 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2009.06753.x
- Primary Citation Related Structures:
3F8N - PubMed Abstract:
In Bacillus subtilis, the transcription factor PerR is an iron dependant sensor of H(2)O(2). The sensing mechanism relies on a selective metal catalysed oxidation of two histidine residues of the regulatory site. Here we present the first crystal structure of the active PerR protein in complex with a Mn(2+) ion. In addition, X-ray absorption spectroscopy experiments were performed to characterize the corresponding iron form of the protein. Both studies reveal a penta-coordinate arrangement of the regulatory site that involves three histidines and two aspartates. One of the histidine ligand belongs to the N-terminal domain. Binding of this residue to the regulatory metal allows the protein to adopt a caliper-like conformation suited to DNA binding. Since this histidine is conserved in all PerR and a vast majority of Fur proteins, it is likely that the allosteric switch induced by the regulatory metal is general for this family of metalloregulators.
- Institut de Biologie Structurale CEA-CNRS-UJF, LCCP, GSY, 41 rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: