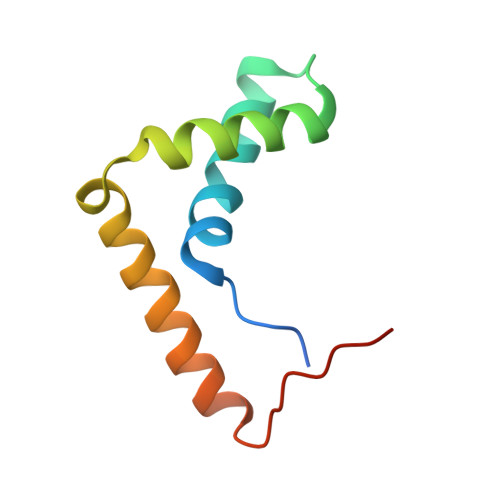
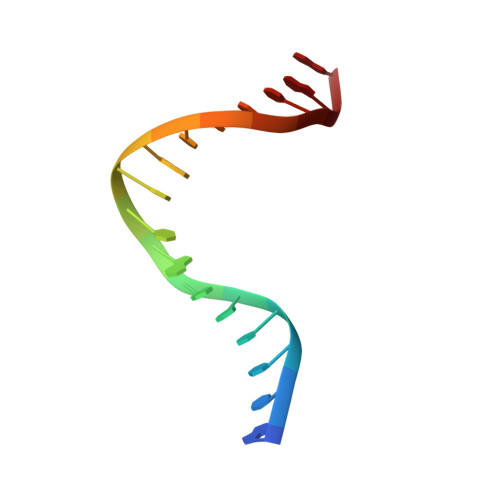
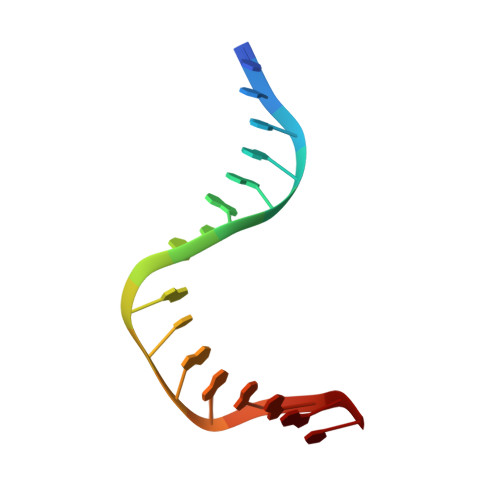
The Structure of Sox17 Bound to DNA Reveals a Conserved Bending Topology but Selective Protein Interaction Platforms
Palasingam, P., Jauch, R., Ng, C.K.L., Kolatkar, P.R.(2009) J Mol Biology 388: 619-630
- PubMed: 19328208 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.055
- Primary Citation Related Structures:
3F27 - PubMed Abstract:
Sox17 regulates endodermal lineage commitment and is thought to function antagonistically to the pluripotency determinant Sox2. To investigate the biochemical basis for the distinct functions of Sox2 and Sox17, we solved the crystal structure of the high mobility group domain of Sox17 bound to a DNA element derived from the Lama1 enhancer using crystals diffracting to 2.7 A resolution. Sox17 targets the minor groove and bends the DNA by approximately 80 degrees . The DNA architecture closely resembles the one seen for Sox2/DNA structures, suggesting that the degree of bending is conserved between both proteins and nucleotide substitutions have only marginal effects on the bending topology. Accordingly, affinities of Sox2 and Sox17 for the Lama1 element were found to be identical. However, when the Oct1 contact interface of Sox2 is compared with the corresponding region of Sox17, a significantly altered charge distribution is observed, suggesting differential co-factor recruitment that may explain their biological distinctiveness.
- Laboratory of Structural Biochemistry, Genome Institute of Singapore, 60 Biopolis Street, Singapore, Singapore.
Organizational Affiliation: