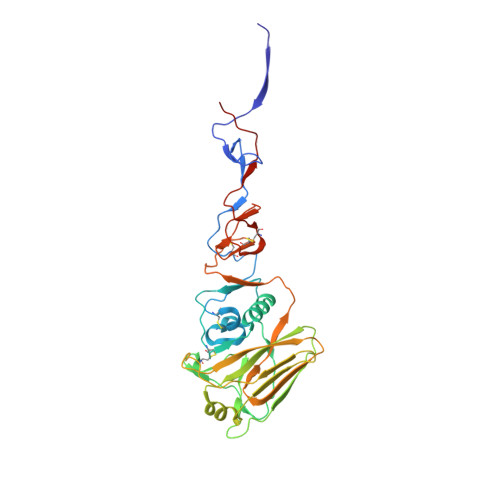
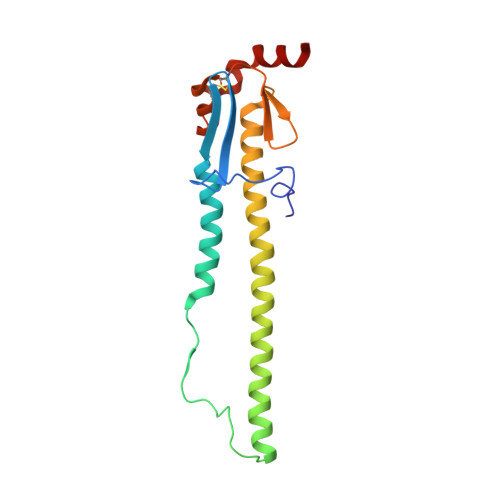
Structure of influenza hemagglutinin in complex with an inhibitor of membrane fusion
Russell, R.J., Kerry, P.S., Stevens, D.J., Steinhauer, D.A., Martin, S.R., Gamblin, S.J., Skehel, J.J.(2008) Proc Natl Acad Sci U S A 105: 17736-17741
- PubMed: 19004788 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0807142105
- Primary Citation Related Structures:
3EYJ, 3EYK, 3EYM - PubMed Abstract:
The influenza surface glycoprotein hemagglutinin (HA) is a potential target for antiviral drugs because of its key roles in the initial stages of infection: receptor binding and the fusion of virus and cell membranes. The structure of HA in complex with a known inhibitor of membrane fusion and virus infectivity, tert-butyl hydroquinone (TBHQ), shows that the inhibitor binds in a hydrophobic pocket formed at an interface between HA monomers. Occupation of this site by TBHQ stabilizes the neutral pH structure through intersubunit and intrasubunit interactions that presumably inhibit the conformational rearrangements required for membrane fusion. The nature of the binding site suggests routes for the chemical modification of TBHQ that could lead to the development of more potent inhibitors of membrane fusion and potential anti-influenza drugs.
- Interdisciplinary Center for Human and Avian Influenza Research, School of Biology, University of St. Andrews, Fife KY16 9ST, United Kingdom. rjmr@st-andrews.ac.uk
Organizational Affiliation: