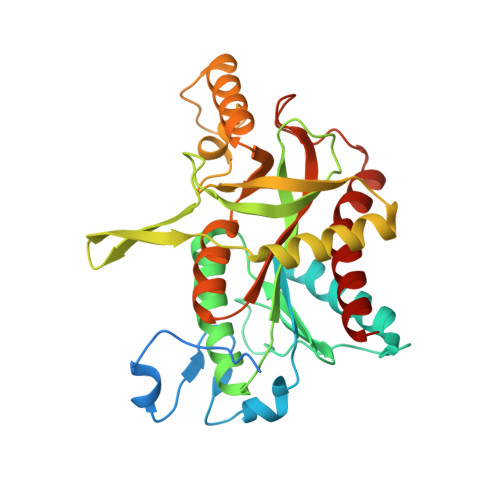
Implications of the structure of human uridine phosphorylase 1 on the development of novel inhibitors for improving the therapeutic window of fluoropyrimidine chemotherapy.
Roosild, T.P., Castronovo, S., Fabbiani, M., Pizzorno, G.(2009) BMC Struct Biol 9: 14-14
- PubMed: 19291308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-9-14
- Primary Citation Related Structures:
3EUE, 3EUF - PubMed Abstract:
Uridine phosphorylase (UPP) is a key enzyme of pyrimidine salvage pathways, catalyzing the reversible phosphorolysis of ribosides of uracil to nucleobases and ribose 1-phosphate. It is also a critical enzyme in the activation of pyrimidine-based chemotherapeutic compounds such a 5-fluorouracil (5-FU) and its prodrug capecitabine. Additionally, an elevated level of this enzyme in certain tumours is believed to contribute to the selectivity of such drugs. However, the clinical effectiveness of these fluoropyrimidine antimetabolites is hampered by their toxicity to normal tissue. In response to this limitation, specific inhibitors of UPP, such as 5-benzylacyclouridine (BAU), have been developed and investigated for their ability to modulate the cytotoxic side effects of 5-FU and its derivatives, so as to increase the therapeutic index of these agents. In this report we present the high resolution structures of human uridine phosphorylase 1 (hUPP1) in ligand-free and BAU-inhibited conformations. The structures confirm the unexpected solution observation that the human enzyme is dimeric in contrast to the hexameric assembly present in microbial UPPs. They also reveal in detail the mechanism by which BAU engages the active site of the protein and subsequently disables the enzyme by locking the protein in a closed conformation. The observed inter-domain motion of the dimeric human enzyme is much greater than that seen in previous UPP structures and may result from the simpler oligomeric organization. The structural details underlying hUPP1's active site and additional surfaces beyond these catalytic residues, which coordinate binding of BAU and other acyclouridine analogues, suggest avenues for future design of more potent inhibitors of this enzyme. Notably, the loop forming the back wall of the substrate binding pocket is conformationally different and substantially less flexible in hUPP1 than in previously studied microbial homologues. These distinctions can be utilized to discover novel inhibitory compounds specifically optimized for efficacy against the human enzyme as a step toward the development of more effective chemotherapeutic regimens that can selectively protect normal tissues with inherently lower UPP activity.
- Department of Drug Development, Nevada Cancer Institute, Las Vegas, Nevada, USA. troosild@nvcancer.org
Organizational Affiliation: