Structure and Functional Studies of the CS Domain of the Essential H/ACA Ribonucleoparticle Assembly Protein SHQ1.
Singh, M., Gonzales, F.A., Cascio, D., Heckmann, N., Chanfreau, G., Feigon, J.(2009) J Biological Chem 284: 1906-1916
- PubMed: 19019820 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M807337200
- Primary Citation Related Structures:
3EUD - PubMed Abstract:
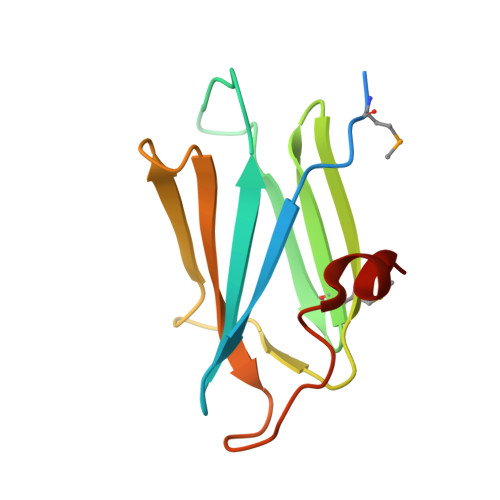
H/ACA ribonucleoprotein particles are essential for ribosomal RNA and telomerase RNA processing and metabolism. Shq1p has been identified as an essential eukaryotic H/ACA small nucleolar (sno) ribonucleoparticle (snoRNP) biogenesis and assembly factor. Shq1p is postulated to be involved in the early biogenesis steps of H/ACA snoRNP complexes, and Shq1p depletion leads to a specific decrease in H/ACA small nucleolar RNA levels and to defects in ribosomal RNA processing. Shq1p contains two predicted domains as follows: an N-terminal CS (named after CHORD-containing proteins and SGT1) or HSP20-like domain, and a C-terminal region of high sequence homology called the Shq1 domain. Here we report the crystal structure and functional studies of the Saccharomyces cerevisiae Shq1p CS domain. The structure consists of a compact anti-parallel beta-sandwich fold that is composed of two beta-sheets containing four and three beta-strands, respectively, and a short alpha-helix. Deletion studies showed that the CS domain is required for the essential functions of Shq1p. Point mutations in residues Phe-6, Gln-10, and Lys-80 destabilize Shq1p in vivo and induce a temperature-sensitive phenotype with depletion of H/ACA small nucleolar RNAs and defects in rRNA processing. Although CS domains are frequently found in co-chaperones of the Hsp90 molecular chaperone, no interaction was detected between the Shq1p CS domain and yeast Hsp90 in vitro. These results show that the CS domain is essential for Shq1p function in H/ACA snoRNP biogenesis in vivo, possibly in an Hsp90-independent manner.
- Department of Chemistry and Biochemistry, UCLA, Los Angeles, California 90095, USA.
Organizational Affiliation: