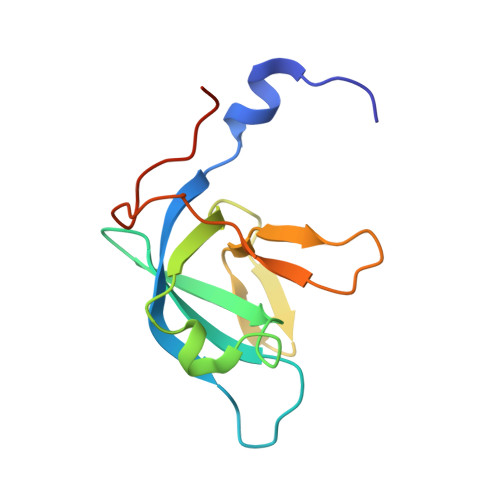
Crystal structure of Trbp111: a tructure specific tRNA binding protein
Swairjo, M.A., Morales, A.J., Wang, C.-C., Ortiz, A.R., Schimmel, P.(2000) EMBO J 19: 6287-6298
- PubMed: 11101501
- DOI: https://doi.org/10.1093/emboj/19.23.6287
- Primary Citation Related Structures:
1PYB, 3ERS - PubMed Abstract:
Trbp111 is a 111 amino acid Aquifex aeolicus structure-specific tRNA-binding protein that has homologous counterparts distributed throughout evolution. A dimer is the functional unit for binding a single tRNA. Here we report the 3D structures of the A.aeolicus protein and its Escherichia coli homolog at resolutions of 2.50 and 1.87 A, respectively. The structure shows a symmetrical dimer of two core domains and a central dimerization domain where the N- and C-terminal regions of Trbp111 form an extensive dimer interface. The core of the monomer is a classical oligonucleotide/oligosaccharide-binding (OB) fold with a five-stranded ss-barrel and a small capping helix. This structure is similar to that seen in the anticodon-binding domain of three class II tRNA synthetases and several other proteins. Mutational analysis identified sites important for interactions with tRNA. These residues line the inner surfaces of two clefts formed between the ss-barrel of each monomer and the dimer interface. The results are consistent with a proposed model for asymmetrical docking of the convex side of tRNA to the dimer.
- Skaags Institute for Chemical Biology, Department of Molecular Biology and Department of Chemistry, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: