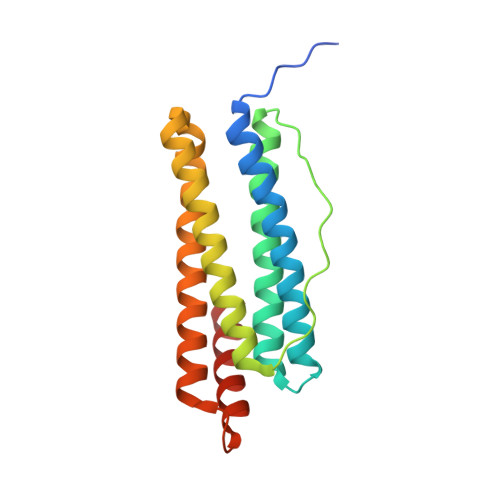
The crystal structure of ferritin from Helicobacter pylori reveals unusual conformational changes for iron uptake.
Cho, K.J., Shin, H.J., Lee, J.H., Kim, K.J., Park, S.S., Lee, Y., Lee, C., Park, S.S., Kim, K.H.(2009) J Mol Biology 390: 83-98
- PubMed: 19427319 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.04.078
- Primary Citation Related Structures:
3EGM - PubMed Abstract:
The crystal structure of recombinant ferritin from Helicobacter pylori has been determined in its apo, low-iron-bound, intermediate, and high-iron-bound states. Similar to other members of the ferritin family, the bacterial ferritin assembles as a spherical protein shell of 24 subunits, each of which folds into a four-alpha-helix bundle. Significant conformational changes were observed at the BC loop and the entrance of the 4-fold symmetry channel in the intermediate and high-iron-bound states, whereas no change was found in the apo and low-iron-bound states. The imidazole rings of His149 at the channel entrance undergo conformational changes that bear resemblance to heme configuration and are directly coupled to axial translocation of Fe ions through the 4-fold channel. Our results provide the first structural evidence of the translocation of Fe ions through the 4-fold channel in prokaryotes and the transition from a protein-dominated process to a mineral-surface-dominated process during biomineralization.
- Department of Life Sciences and Biotechnology, Korea University, Seoul, Korea.
Organizational Affiliation: