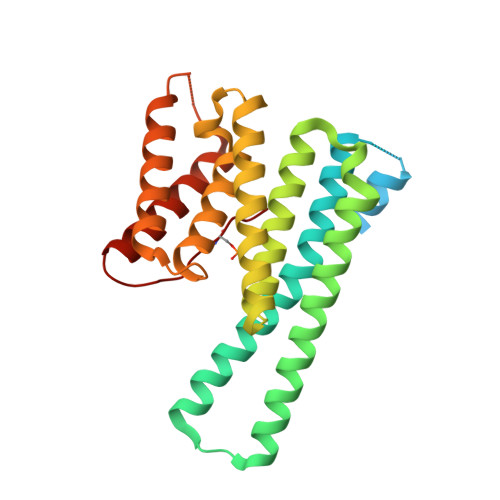
Characterization of 14-3-3 proteins from Cryptosporidium parvum.
Brokx, S.J., Wernimont, A.K., Dong, A., Wasney, G.A., Lin, Y.H., Lew, J., Vedadi, M., Lee, W.H., Hui, R.(2011) PLoS One 6: e14827-e14827
- PubMed: 21853016 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0014827
- Primary Citation Related Structures:
2NPM, 2O8P, 3EFZ - PubMed Abstract:
The parasite Cryptosporidium parvum has three 14-3-3 proteins: Cp14ε, Cp14a and Cp14b, with only Cp14ε similar to human 14-3-3 proteins in sequence, peptide-binding properties and structure. Structurally, Cp14a features the classical 14-3-3 dimer but with a uniquely wide pocket and a disoriented RRY triad potentially incapable of binding phosphopeptides. The Cp14b protein deviates from the norm significantly: (i) In one subunit, the phosphorylated C-terminal tail is bound in the binding groove like a phosphopeptide. This supports our binding study indicating this protein was stabilized by a peptide mimicking its last six residues. (ii) The other subunit has eight helices instead of nine, with αA and αB forming a single helix and occluding the peptide-binding cleft. (iii) The protein forms a degenerate dimer with the two binding grooves divided and facing opposite directions. These features conspire to block and disrupt the bicameral substrate-binding pocket, suggesting a possible tripartite auto-regulation mechanism that has not been observed previously. This article can also be viewed as an enhanced version in which the text of the article is integrated with interactive 3D representations and animated transitions. Please note that a web plugin is required to access this enhanced functionality. Instructions for the installation and use of the web plugin are available in Text S1.
- Structural Genomics Consortium, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: