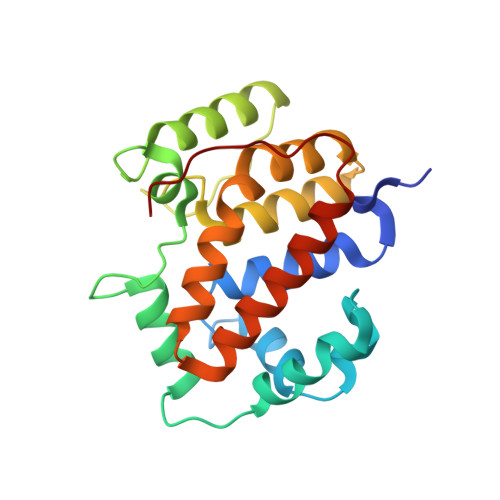
Structural analysis of a monomeric form of the twin-arginine leader peptide binding chaperone Escherichia coli DmsD.
Stevens, C.M., Winstone, T.M., Turner, R.J., Paetzel, M.(2009) J Mol Biology 389: 124-133
- PubMed: 19361518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.03.069
- Primary Citation Related Structures:
3EFP - PubMed Abstract:
The redox enzyme maturation proteins play an essential role in the proofreading and membrane targeting of protein substrates to the twin-arginine translocase. Functionally, the most thoroughly characterized redox enzyme maturation protein to date is Escherichia coli DmsD (EcDmsD). Herein, we present the X-ray crystal structure of the monomeric form of the EcDmsD refined to 2.0 A resolution, with clear electron density present for each of its 204 amino acid residues. The structural data presented here complement the biochemical data previously generated regarding the function of these twin-arginine translocase leader peptide binding chaperone proteins. Docking and molecular dynamics simulation experiments were used to provide a proposed model for how this chaperone is able to recognize the leader peptide of its substrate DmsA. The interactions observed in the model are in agreement with previous biochemical data and suggest intimate interactions between the conserved twin-arginine motif of the leader peptide of E. coli DmsA and the most conserved regions on the surface of EcDmsD.
- Department of Molecular Biology and Biochemistry, Simon Fraser University, Burnaby, British Columbia, Canada.
Organizational Affiliation: