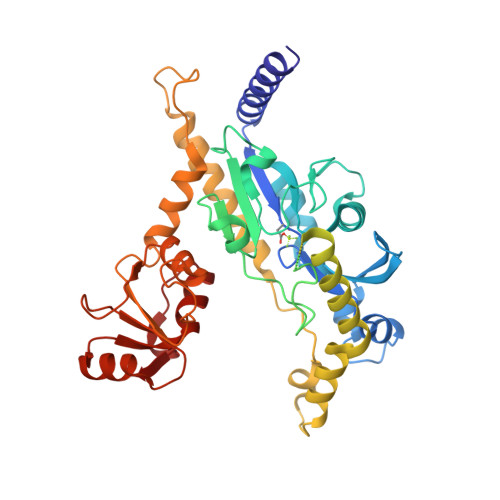
The structure of Fcp1, an essential RNA polymerase II CTD phosphatase.
Ghosh, A., Shuman, S., Lima, C.D.(2008) Mol Cell 32: 478-490
- PubMed: 19026779 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2008.09.021
- Primary Citation Related Structures:
3EF0, 3EF1 - PubMed Abstract:
Kinases and phosphatases regulate mRNA synthesis and processing by phosphorylating and dephosphorylating the C-terminal domain (CTD) of the largest subunit of RNA polymerase II. Fcp1 is an essential CTD phosphatase that preferentially hydrolyzes Ser2-PO(4) of the tandem YSPTSPS CTD heptad array. Fcp1 crystal structures were captured at two stages of the reaction pathway: a Mg-BeF(3) complex that mimics the aspartylphosphate intermediate and a Mg-AlF(4)(-) complex that mimics the transition state of the hydrolysis step. Fcp1 is a Y-shaped protein composed of an acylphosphatase domain located at the base of a deep canyon formed by flanking modules that are missing from the small CTD phosphatase (SCP) clade: an Fcp1-specific helical domain and a C-terminal BRCA1 C-terminal (BRCT) domain. The structure and mutational analysis reveals that Fcp1 and Scp1 (a Ser5-selective phosphatase) adopt different CTD-binding modes; we surmise the CTD threads through the Fcp1 canyon to access the active site.
- Structural Biology Program, Sloan-Kettering Institute, New York, NY 10021, USA.
Organizational Affiliation: