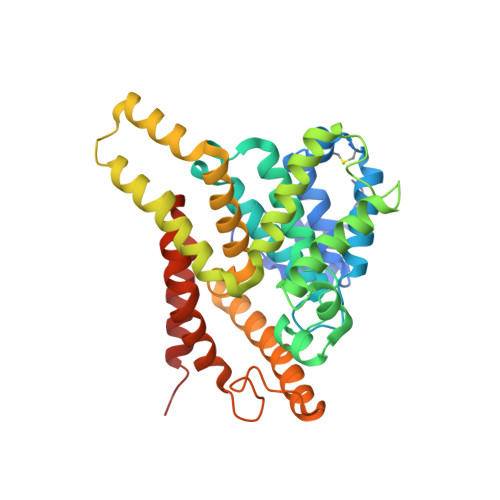
Kinetic and structural studies of phosphodiesterase-8A and implication on the inhibitor selectivity
Wang, H., Yan, Z., Yang, S., Cai, J., Robinson, H., Ke, H.(2008) Biochemistry 47: 12760-12768
- PubMed: 18983167 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi801487x
- Primary Citation Related Structures:
3ECM, 3ECN - PubMed Abstract:
Cyclic nucleotide phosphodiesterase-8 (PDE8) is a family of cAMP-specific enzymes and plays important roles in many biological processes, including T-cell activation, testosterone production, adrenocortical hyperplasia, and thyroid function. However, no PDE8 selective inhibitors are available for trial treatment of human diseases. Here we report kinetic properties of the highly active PDE8A1 catalytic domain prepared from refolding and its crystal structures in the unliganded and 3-isobutyl-1-methylxanthine (IBMX) bound forms at 1.9 and 2.1 A resolutions, respectively. The PDE8A1 catalytic domain has a K(M) of 1.8 microM, V(max) of 6.1 micromol/min/mg, a k(cat) of 4.0 s(-1) for cAMP, and a K(M) of 1.6 mM, V(max) of 2.5 micromol/min/mg, a k(cat) of 1.6 s(-1) for cGMP, thus indicating that the substrate specificity of PDE8 is dominated by K(M). The structure of the PDE8A1 catalytic domain has similar topology as those of other PDE families but contains two extra helices around Asn685-Thr710. Since this fragment is distant from the active site of the enzyme, its impact on the catalysis is unclear. The PDE8A1 catalytic domain is insensitive to the IBMX inhibition (IC(50) = 700 microM). The unfavorable interaction of IBMX in the PDE8A1-IBMX structure suggests an important role of Tyr748 in the inhibitor binding. Indeed, the mutation of Tyr748 to phenylalanine increases the PDE8A1 sensitivity to several nonselective or family selective PDE inhibitors. Thus, the structural and mutagenesis studies provide not only insight into the enzymatic properties but also guidelines for design of PDE8 selective inhibitors.
- Department of Biochemistry and Biophysics, Lineberger Comprehensive Cancer Center, The University of North Carolina, Chapel Hill, North Carolina 27599-7260, USA.
Organizational Affiliation: