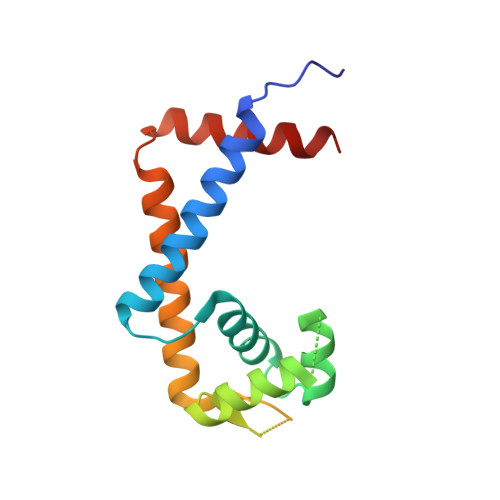
The crystal structure of MexR from Pseudomonas aeruginosa in complex with its antirepressor ArmR
Wilke, M.S., Heller, M., Creagh, A.L., Haynes, C.A., McIntosh, L.P., Poole, K., Strynadka, N.C.J.(2008) Proc Natl Acad Sci U S A 105: 14832-14837
- PubMed: 18812515 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0805489105
- Primary Citation Related Structures:
3ECH - PubMed Abstract:
The intrinsic antimicrobial resistance of the opportunistic human pathogen Pseudomonas aeruginosa is compounded in mutant strains that overexpress multidrug efflux pumps such as the prominent drug-proton antiporter, MexAB-OprM. The primary regulator of the mexAB-oprM operon is the MarR family repressor, MexR. An additional repressor, NalC, also regulates mexAB-oprM by controlling expression of ArmR, an antirepressor peptide that is hypothesized to prevent the binding of MexR to its cognate DNA operator via an allosteric protein-peptide interaction. To better understand how ArmR modulates MexR, we determined the MexR-binding region of ArmR as its C-terminal 25 residues and solved the crystal structure of MexR in a 2:1 complex with this ArmR fragment at 1.8 A resolution. This structure reveals that the C-terminal residues of ArmR form a kinked alpha-helix, which occupies a pseudosymmetrical and largely hydrophobic binding cavity located at the centre of the MexR dimer. Although the ArmR-binding cavity partially overlaps with the small molecule effector-binding sites of other MarR family members, it possesses a larger and more complex binding surface to accommodate the greater size and specific physicochemical properties of a peptide effector. Comparison with the structure of apo-MexR reveals that ArmR stabilizes a dramatic conformational change that is incompatible with DNA-binding. Thus, this work defines the structural mechanism by which ArmR allosterically derepresses MexR-controlled gene expression in P. aeruginosa and reveals important insights into the regulation of multidrug resistance.
- Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, BC, Canada V6T1Z3.
Organizational Affiliation: