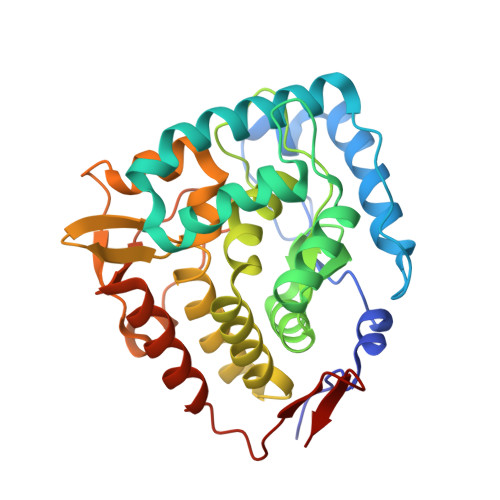
Crystal structure of tryptophan hydroxylase with bound amino acid substrate
Windahl, M.S., Petersen, C.R., Christensen, H.E.M., Harris, P.(2008) Biochemistry 47: 12087-12094
- PubMed: 18937498 Search on PubMed
- DOI: https://doi.org/10.1021/bi8015263
- Primary Citation Related Structures:
3E2T - PubMed Abstract:
Tryptophan hydroxylase (TPH) is a mononuclear non-heme iron enzyme, which catalyzes the reaction between tryptophan, O 2, and tetrahydrobiopterin (BH 4) to produce 5-hydroxytryptophan and 4a-hydroxytetrahydrobiopterin. This is the first and rate-limiting step in the biosynthesis of the neurotransmitter and hormone serotonin (5-hydroxytryptamine). We have determined the 1.9 A resolution crystal structure of the catalytic domain (Delta1-100/Delta415-445) of chicken TPH isoform 1 (TPH1) in complex with the tryptophan substrate and an iron-bound imidazole. This is the first structure of any aromatic amino acid hydroxylase with bound natural amino acid substrate. The iron coordination can be described as distorted trigonal bipyramidal coordination with His273, His278, and Glu318 (partially bidentate) and one imidazole as ligands. The tryptophan stacks against Pro269 with a distance of 3.9 A between the iron and the tryptophan Czeta3 atom that is hydroxylated. The binding of tryptophan and maybe the imidazole has caused the structural changes in the catalytic domain compared to the structure of the human TPH1 without tryptophan. The structure of chicken TPH1 is more compact, and the loops of residues Leu124-Asp139 and Ile367-Thr369 close around the active site. Similar structural changes are seen in the catalytic domain of phenylalanine hydroxylase (PAH) upon binding of substrate analogues norleucine and thienylalanine to the PAH.BH 4 complex. In fact, the chicken TPH1.Trp.imidazole structure resembles the PAH.BH 4.thienylalanine structure more (root-mean-square deviation for Calpha atoms of 0.90 A) than the human TPH1 structure (root-mean-square deviation of 1.47 A).
- Department of Basic Sciences and Environment, Faculty of Life Sciences, University of Copenhagen, Thorvaldsensvej 40, 1871 Frederiksberg C, Denmark.
Organizational Affiliation: