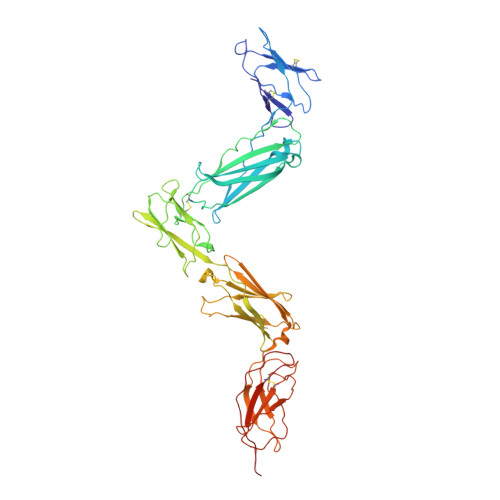
Structural organization of a full-length gp130/LIF-R cytokine receptor transmembrane complex.
Skiniotis, G., Lupardus, P.J., Martick, M., Walz, T., Garcia, K.C.(2008) Mol Cell 31: 737-748
- PubMed: 18775332 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2008.08.011
- Primary Citation Related Structures:
3E0G - PubMed Abstract:
gp130 is a shared receptor for at least nine cytokines and can signal either as a homodimer or as a heterodimer with Leukemia Inhibitory Factor Receptor (LIF-R). Here, we biophysically and structurally characterize the full-length, transmembrane form of a quaternary cytokine receptor complex consisting of gp130, LIF-R, the cytokine Ciliary Neurotrophic Factor (CNTF), and its alpha receptor (CNTF-Ralpha). Thermodynamic analysis indicates that, unlike the cooperative assembly of the symmetric gp130/Interleukin-6/IL-6Ralpha hexameric complex, CNTF/CNTF-Ralpha heterodimerizes gp130 and LIF-R via noncooperative energetics to form an asymmetric 1:1:1:1 complex. Single particle electron microscopic analysis of the full-length gp130/LIF-R/CNTF-Ralpha/CNTF quaternary complex elucidates an asymmetric structural arrangement, in which the receptor extracellular and transmembrane segments join as a continuous, rigid unit, poised to sensitively transduce ligand engagement to the membrane-proximal intracellular signaling regions. These studies also enumerate the organizing principles for assembly of the "tall" class of gp130 family cytokine receptor complexes including LIF, IL-27, IL-12, and others.
- Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115, USA.
Organizational Affiliation: