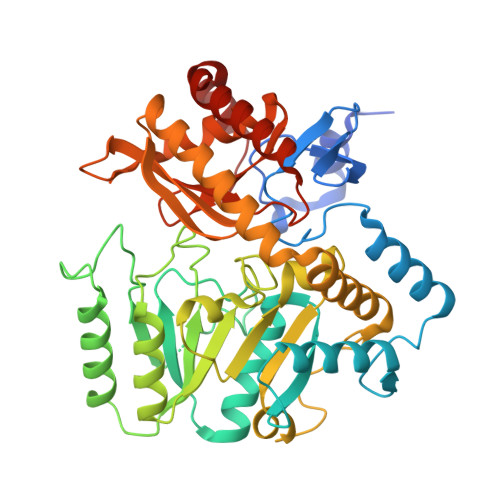
The novel structure of a pyridoxal 5'-phosphate-dependent fold-type I racemase, alpha-amino-epsilon-caprolactam racemase from Achromobacter obae
Okazaki, S., Suzuki, A., Mizushima, T., Kawano, T., Komeda, H., Asano, Y., Yamane, T.(2009) Biochemistry 48: 941-950
- PubMed: 19146406 Search on PubMed
- DOI: https://doi.org/10.1021/bi801574p
- Primary Citation Related Structures:
2ZUK, 3DXV, 3DXW - PubMed Abstract:
Alpha-amino-epsilon-caprolactam (ACL) racemase (ACLR) from Achromobacter obae catalyzes the interconversion of l- and d-ACL. ACLR belongs to the fold-type I group of pyridoxal 5'-phosphate (PLP) dependent enzymes. In this study, the first crystal structures of a fold-type I racemase are solved for the native form and epsilon-caprolactam-complexed form of ACLR at 2.21 and 2.40 A resolution, respectively. Based on the location of epsilon-caprolactam in the complex structure, the substrate-binding site is assigned between Trp49 and Tyr137. The carboxyl group of Asp210 is a reasonable candidate that recognizes the nitrogen atom of a lactam or amide in the substrate. Based on a structural comparison with fold-type III alanine racemase, Tyr137 is potentially the acid/base catalytic residue that is essential for the two-base racemization mechanism. The overall structure of ACLR is similar to that of fold-type I enzymes. A structural comparison with these enzymes explains the different reaction specificities.
- Department of Biotechnology, Graduate School of Engineering, Nagoya University, Chikusa, Nagoya 464-8603, Japan.
Organizational Affiliation: