The RNA polymerase "switch region" is a target for inhibitors.
Mukhopadhyay, J., Das, K., Ismail, S., Koppstein, D., Jang, M., Hudson, B., Sarafianos, S., Tuske, S., Patel, J., Jansen, R., Irschik, H., Arnold, E., Ebright, R.H.(2008) Cell 135: 295-307
- PubMed: 18957204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2008.09.033
- Primary Citation Related Structures:
3DXJ - PubMed Abstract:
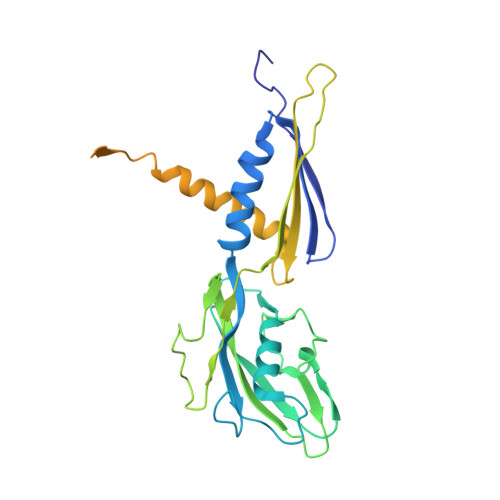
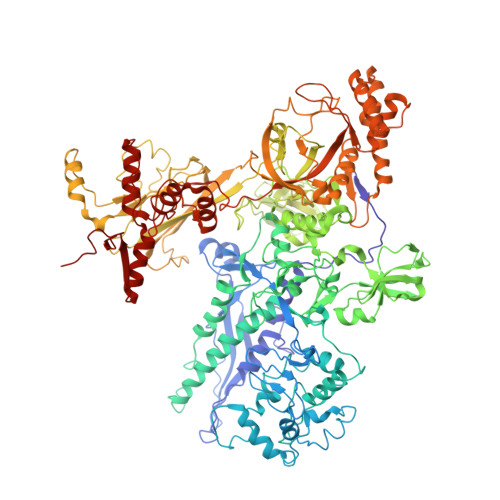
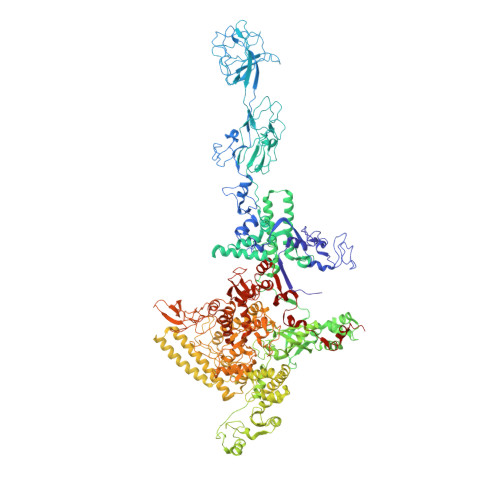
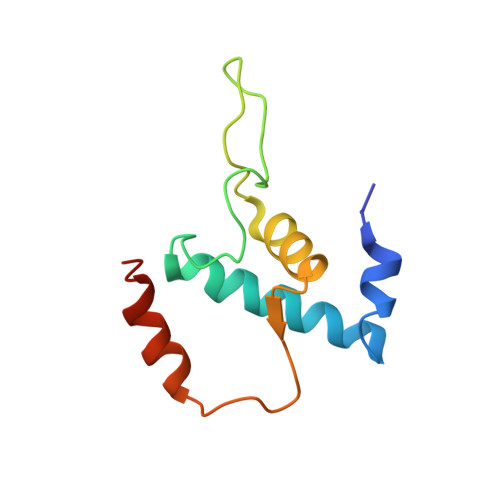
The alpha-pyrone antibiotic myxopyronin (Myx) inhibits bacterial RNA polymerase (RNAP). Here, through a combination of genetic, biochemical, and structural approaches, we show that Myx interacts with the RNAP "switch region"--the hinge that mediates opening and closing of the RNAP active center cleft--to prevent interaction of RNAP with promoter DNA. We define the contacts between Myx and RNAP and the effects of Myx on RNAP conformation and propose that Myx functions by interfering with opening of the RNAP active-center cleft during transcription initiation. We further show that the structurally related alpha-pyrone antibiotic corallopyronin (Cor) and the structurally unrelated macrocyclic-lactone antibiotic ripostatin (Rip) function analogously to Myx. The RNAP switch region is distant from targets of previously characterized RNAP inhibitors, and, correspondingly, Myx, Cor, and Rip do not exhibit crossresistance with previously characterized RNAP inhibitors. The RNAP switch region is an attractive target for identification of new broad-spectrum antibacterial therapeutic agents.
- Howard Hughes Medical Institute, Rutgers University, Piscataway, NJ 08854, USA.
Organizational Affiliation: