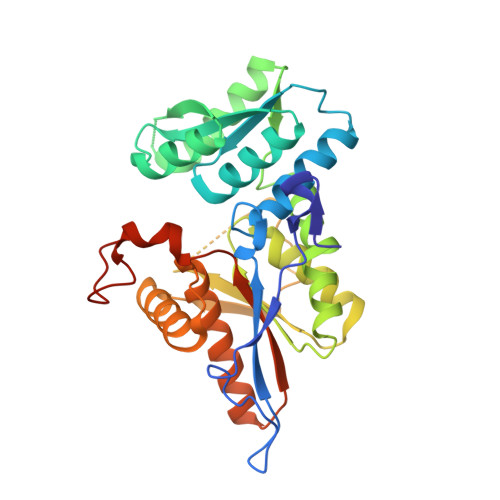
Crystal structure of a sulfur carrier protein complex found in the cysteine biosynthetic pathway of Mycobacterium tuberculosis.
Jurgenson, C.T., Burns, K.E., Begley, T.P., Ealick, S.E.(2008) Biochemistry 47: 10354-10364
- PubMed: 18771296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi800915j
- Primary Citation Related Structures:
3DWG, 3DWI, 3DWM - PubMed Abstract:
The structure of the protein complex CysM-CysO from a new cysteine biosynthetic pathway found in the H37Rv strain of Mycobacterium tuberculosis has been determined at 1.53 A resolution. CysM (Rv1336) is a PLP-containing beta-replacement enzyme and CysO (Rv1335) is a sulfur carrier protein with a ubiquitin-like fold. CysM catalyzes the replacement of the acetyl group of O-acetylserine by CysO thiocarboxylate to generate a protein-bound cysteine that is released in a subsequent proteolysis reaction. The protein complex in the crystal structure is asymmetric with one CysO protomer binding to one end of a CysM dimer. Additionally, the structures of CysM and CysO were determined individually at 2.8 and 2.7 A resolution, respectively. Sequence alignments with homologues and structural comparisons with CysK, a cysteine synthase that does not utilize a sulfur carrier protein, revealed high conservation of active site residues; however, residues in CysM responsible for CysO binding are not conserved. Comparison of the CysM-CysO binding interface with other sulfur carrier protein complexes revealed a similarity in secondary structural elements that contribute to complex formation in the ThiF-ThiS and MoeB-MoaD systems, despite major differences in overall folds. Comparison of CysM with and without bound CysO revealed conformational changes associated with CysO binding.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853-1301, USA.
Organizational Affiliation: