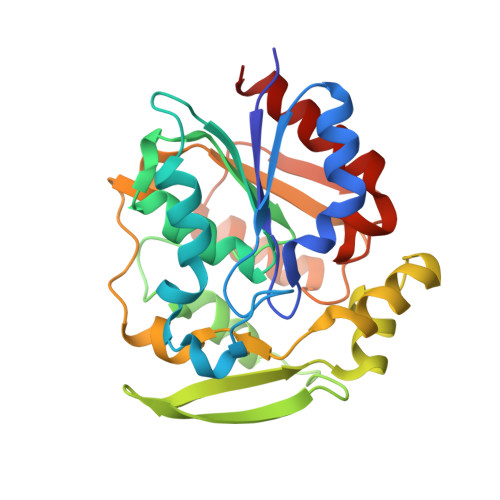
Hydroxynitrile lyases with alpha / beta-hydrolase fold: two enzymes with almost identical 3D structures but opposite enantioselectivities and different reaction mechanisms
Andexer, J.N., Staunig, N., Eggert, T., Kratky, C., Pohl, M., Gruber, K.(2012) Chembiochem 13: 1932-1939
- PubMed: 22851196
- DOI: https://doi.org/10.1002/cbic.201200239
- Primary Citation Related Structures:
3DQZ - PubMed Abstract:
Hydroxynitrile lyases (HNLs) catalyze the cleavage of cyanohydrins to yield hydrocyanic acid (HCN) and the respective carbonyl compound and are key enzymes in the process of cyanogenesis in plants. In organic syntheses, HNLs are used as biocatalysts for the formation of enantiopure cyanohydrins. We determined the structure of the recently identified, R-selective HNL from Arabidopsis thaliana (AtHNL) at a crystallographic resolution of 2.5 Å. The structure exhibits an α/β-hydrolase fold, very similar to the homologous, but S-selective, HNL from Hevea brasiliensis (HbHNL). The similarities also extend to the active sites of these enzymes, with a Ser-His-Asp catalytic triad present in all three cases. In order to elucidate the mode of substrate binding and to understand the unexpected opposite enantioselectivity of AtHNL, complexes of the enzyme with both (R)- and (S)-mandelonitrile were modeled using molecular docking simulations. Compared to the complex of HbHNL with (S)-mandelonitrile, the calculations produced an approximate mirror image binding mode of the substrate with the phenyl rings located at very similar positions, but with the cyano groups pointing in opposite directions. A catalytic mechanism for AtHNL is proposed, in which His236 from the catalytic triad acts as a general base and the emerging negative charge on the cyano group is stabilized by main-chain amide groups and an α-helix dipole very similar to α/β-hydrolases. This mechanistic proposal is additionally supported by mutagenesis studies.
- Institute of Pharmaceutical Sciences, Albert-Ludwigs-University Freiburg, Albertstrasse 25, Freiburg, Germany.
Organizational Affiliation: